Hi again,
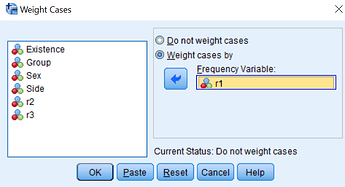
The data are aggregated data and this is why it needs weighting.
temp <- structure(list(Existence = structure(c(2L, 1L, 2L, 1L, 2L, 1L,
2L, 1L, 2L, 1L, 2L, 1L, 2L, 1L, 2L, 1L), .Label = c("not_exists",
"exists"), class = "factor"), Group = structure(c(2L, 2L, 1L,
1L, 2L, 2L, 1L, 1L, 2L, 2L, 1L, 1L, 2L, 2L, 1L, 1L), .Label = c("Group_1",
"Group_2"), class = "factor"), Sex = structure(c(1L, 1L, 1L,
1L, 1L, 1L, 1L, 1L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L), .Label = c("F",
"M"), class = "factor"), Side = structure(c(1L, 1L, 1L, 1L, 2L,
2L, 2L, 2L, 1L, 1L, 1L, 1L, 2L, 2L, 2L, 2L), .Label = c("right",
"left"), class = "factor"), r1 = c(35, 77, 28, 70, 37, 75, 24,
74, 23, 27, 21, 31, 20, 30, 18, 34), r2 = c(21, 91, 7, 91, 17,
95, 8, 90, 18, 32, 5, 47, 19, 31, 7, 45), r3 = c(62, 50, 41,
57, 47, 65, 35, 63, 28, 22, 29, 23, 28, 22, 31, 21)), class = "data.frame", row.names = c(NA,
-16L), variable.labels = structure(character(0), .Names = character(0)), codepage = 65001L)
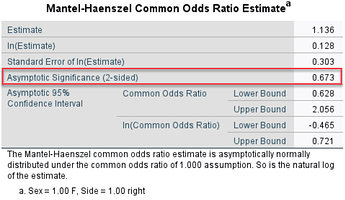
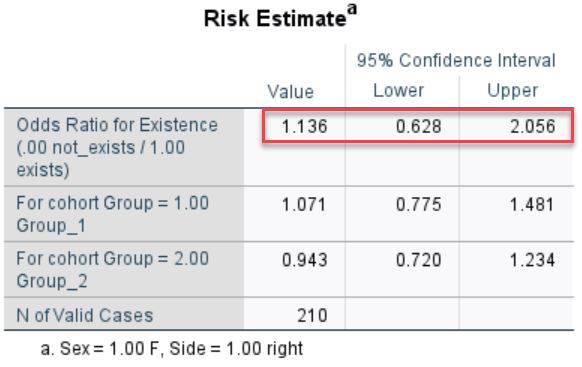
I would like to calculate Odds Ratio (OR) and confidence intervals for it and p-value by means of firstly weighting cases separately, I think by r1, r2, r3 variables. I have read what SPSS does is weighting by frequency if I am not mistaken.
The results I would like to place in dataframe.
For example the results calculated in SPSS ( for OR, CI, p-value) comparing Group_1 and Group_2 for Female on the right hand side are as follows: