I'm unable to run the predict function on my fit object.
Here is my r code :
#load libraries
library(tidymodels)
library(tidyverse)
#using mtcars as example
thisdf <- mtcars
response.var <- 'mpg'
#eliminate non-numeric columns
thisdf2 <- thisdf %>%
select_if(~is.numeric(.x))
#eliminate rows with missing response variables
thisdf2 <- thisdf2[!is.na(thisdf2[,response.var]),]
#train test split
thisdf2.split <- initial_split(thisdf2, prop=0.6)
thisdf2.train <- training (thisdf2.split)
thisdf2.test <- testing (thisdf2.split)
#formula
f <- as.formula(paste0(response.var, ' ~ .'))
#5 fold cross validation
thisdf.cv <- vfold_cv(thisdf2.train, v=5)
#recipe
this.recipe <- recipe(f, data=thisdf2.train) %>%
step_corr(all_predictors()) %>%
step_nzv(all_predictors()) %>%
step_medianimpute(all_predictors()) %>%
step_center(all_predictors()) %>%
step_scale(all_predictors())
this.model <- rand_forest(mode='regression') %>% #,mtry=tune(), trees=tune()
set_engine("ranger", importance='impurity')
this.wf <- workflow() %>%
add_model(this.model) %>%
add_recipe(this.recipe)
train.wf <- this.wf %>%
fit(thisdf2.train)
#measure performance
thisdf2.test$preds <- predict(train.wf, newdata = thisdf2.test)
The predict call throws the following error :
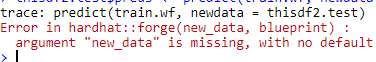
I tried using skip=TRUE argument for the recipe arguments but still get the error.