HI @Slavek, Welcome to RStudio Community!
What have you tried so far? It will be much easier to help you if you turn this into a self-contained reprex (short for reproducible example)? It will help us help you if we can be sure we're all working with/looking at the same stuff.
install.packages("reprex")
If you've never heard of a reprex before, you might want to start by reading the tidyverse.org help page. The reprex dos and don'ts are also useful.
What to do if you run into clipboard problems
If you run into problems with access to your clipboard, you can specify an outfile for the reprex, and then copy and paste the contents into the forum.
reprex::reprex(input = "fruits_stringdist.R", outfile = "fruits_stringdist.md")
For pointers specific to the community site, check out the reprex FAQ, linked to below.
If you simply want to create a new column with the combined products you can do this:
library(tibble)
library(dplyr)
#>
#> Attaching package: 'dplyr'
#> The following objects are masked from 'package:stats':
#>
#> filter, lag
#> The following objects are masked from 'package:base':
#>
#> intersect, setdiff, setequal, union
library(stringr)
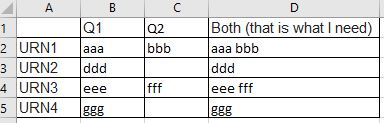
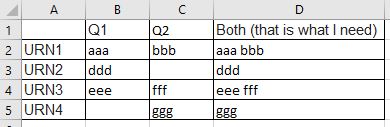
my_df <- tribble(
~name, ~q1, ~q2,
"URN1", "aaa", "bbb",
"URN2", "ddd", NA_character_,
"URN3", "eee", "fff",
"URN4", NA_character_, "ggg"
)
my_df %>%
mutate_at(vars(q1, q2), funs(str_replace_na(., replacement = ""))) %>%
mutate(both = paste(q1, q2) %>% str_trim())
#> # A tibble: 4 x 4
#> name q1 q2 both
#> <chr> <chr> <chr> <chr>
#> 1 URN1 aaa bbb aaa bbb
#> 2 URN2 ddd "" ddd
#> 3 URN3 eee fff eee fff
#> 4 URN4 "" ggg ggg
Created on 2018-10-03 by the reprex package (v0.2.0).
This example assumes that the empty strings are NA values. If that is not that case then you can skip the mutate_at line