Hello,
I have a large dataframe (1000 rows and 200 columns) consisting of several values (names of something ). I subset and retain only rows that have certain values of my choice (eg football, camera, car,plane)
Thanks
Hello,
I have a large dataframe (1000 rows and 200 columns) consisting of several values (names of something ). I subset and retain only rows that have certain values of my choice (eg football, camera, car,plane)
Thanks
Take a look at the filter function in the package dplyr. Subset rows using column values — filter • dplyr
Here's a tiny example of how you would do this but if you need more help. Please look into providing a reproducible example (FAQ: How to do a minimal reproducible example ( reprex ) for beginners):
library(tidyverse)
subsetteddata <- dataset %>%
filter(variablename %in% c("football", "camera", "car", "plane"))
Thanks, it still does not give me what I need. Maybe I will need to explain my self well.
I have a dataframe, and in reality it has 50 variables (columns)- patients diagnosis (I10, A43, Y66, R54, Y67, E32...). I want to extract all patients (rows) with certain diagnosis (I10, A43, Y66). Any of those diagnosis can be in any column but they are mutually exclusive.
I think you need to provide a reproducible example at this point so we can best help you.
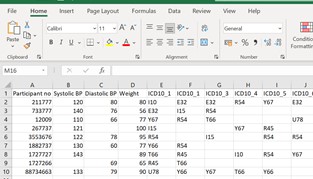
Please see an extract from the df. I need to delete any value from the ICD_10 columns that is not either I10 or Y67 or E32- Then, extract/create a new df which includes only rows with either I10 or Y67 or E32.
This isn't a reproducible example. We can't do much with a screenshot. Please read this and see how to set up a reproducible example.
library(tidyverse)
df <- tibble(
name = rep(seq(1:10), 100),
id = 1 + floor(0.1*(0:(length(name)-1))),
value = (LETTERS)[ntile(runif(1000), 26)]
) %>%
pivot_wider(names_prefix = "IDC_")
df
#> # A tibble: 100 x 11
#> id IDC_1 IDC_2 IDC_3 IDC_4 IDC_5 IDC_6 IDC_7 IDC_8 IDC_9 IDC_10
#> <dbl> <chr> <chr> <chr> <chr> <chr> <chr> <chr> <chr> <chr> <chr>
#> 1 1 E Y Q D S C K Q Z O
#> 2 2 B V B L C B B U E E
#> 3 3 F G C T S G K T T W
#> 4 4 Q N C Z B F W B U B
#> 5 5 G S X J S N N W I H
#> 6 6 J C J N K C U H X M
#> 7 7 O T Q P P I P I D O
#> 8 8 K I S Z R F Z R Y B
#> 9 9 Y P I R U M C N C N
#> 10 10 B P G P A J V V S O
#> # ... with 90 more rows
df_deleted <- df %>%
pivot_longer(starts_with("IDC")) %>%
mutate(value = if_else(
value %in% c("A", "B", "C"),
NA_character_,
value
)) %>%
pivot_wider()
df_deleted
#> # A tibble: 100 x 11
#> id IDC_1 IDC_2 IDC_3 IDC_4 IDC_5 IDC_6 IDC_7 IDC_8 IDC_9 IDC_10
#> <dbl> <chr> <chr> <chr> <chr> <chr> <chr> <chr> <chr> <chr> <chr>
#> 1 1 E Y Q D S <NA> K Q Z O
#> 2 2 <NA> V <NA> L <NA> <NA> <NA> U E E
#> 3 3 F G <NA> T S G K T T W
#> 4 4 Q N <NA> Z <NA> F W <NA> U <NA>
#> 5 5 G S X J S N N W I H
#> 6 6 J <NA> J N K <NA> U H X M
#> 7 7 O T Q P P I P I D O
#> 8 8 K I S Z R F Z R Y <NA>
#> 9 9 Y P I R U M <NA> N <NA> N
#> 10 10 <NA> P G P <NA> J V V S O
#> # ... with 90 more rows
df_only <- df %>%
filter(select(., -id) %>% apply(1, function(x) any(x %in% c("A", "B", "C"))))
df_only
#> # A tibble: 72 x 11
#> id IDC_1 IDC_2 IDC_3 IDC_4 IDC_5 IDC_6 IDC_7 IDC_8 IDC_9 IDC_10
#> <dbl> <chr> <chr> <chr> <chr> <chr> <chr> <chr> <chr> <chr> <chr>
#> 1 1 E Y Q D S C K Q Z O
#> 2 2 B V B L C B B U E E
#> 3 3 F G C T S G K T T W
#> 4 4 Q N C Z B F W B U B
#> 5 6 J C J N K C U H X M
#> 6 8 K I S Z R F Z R Y B
#> 7 9 Y P I R U M C N C N
#> 8 10 B P G P A J V V S O
#> 9 11 N I D C G J S I P D
#> 10 13 A N H X L G J Y V L
#> # ... with 62 more rows
Created on 2021-06-17 by the reprex package (v1.0.0)
The preferred way to do row-wise operations is now rowwise(), but I can never seem to figure out how to use it beyond very simple cases. I find apply(x, 1, fn) much easier to understand.
This topic was automatically closed 21 days after the last reply. New replies are no longer allowed.
If you have a query related to it or one of the replies, start a new topic and refer back with a link.