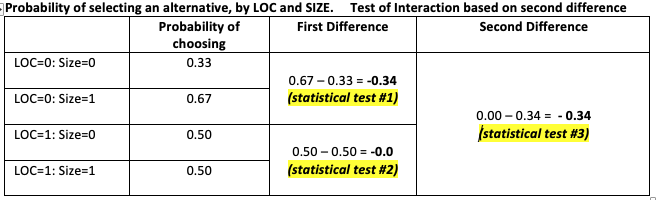
The LOC.SIZE object doesn't contain the p-values
suppressPackageStartupMessages({library(ggeffects)
library(mlogit)
library(sjPlot)})
cedata.1 <- data.frame( id = c(1,1,1,1,1,1,2,2,2,2,2,2), # respondent ID.
QES = c(1,1,2,2,3,3,1,1,2,2,3,3), # Choice set (with 2 alternatives)
Alt = c(1,2,1,2,1,2,1,2,1,2,1,2), # Alt 1 or Alt 2 in choice set
LOC = c(0,0,1,1,0,1,0,1,1,0,0,1), # attribute describing alternative. binary categorical variable
SIZE = c(1,1,1,0,0,1,0,0,1,1,0,1), # attribute describing alternative. binary categorical variable
Choice = c(0,1,1,0,1,0,0,1,0,1,0,1), # if alternative is Chosen (1) or not (0)
gender = c(1,1,1,1,1,1,0,0,0,0,0,0) # male or female (repeats for each indivdual)
)
cedata.1$Choice <- as.factor(cedata.1$Choice)
cedata.1$LOC <- as.factor(cedata.1$LOC)
cedata.1$SIZE <- as.factor(cedata.1$SIZE)
glm.model <- glm(Choice ~ LOC*SIZE, data=cedata.1, family = binomial(link = "logit"))
LOC.SIZE <- ggpredict(glm.model, terms = c("LOC", "SIZE"))
str(LOC.SIZE)
#> Classes 'ggeffects' and 'data.frame': 4 obs. of 6 variables:
#> $ x : Factor w/ 2 levels "0","1": 1 1 2 2
#> $ predicted: num 0.333 0.667 0.5 0.5
#> $ std.error: num 1.22 1.22 1.41 1
#> $ conf.low : num 0.0434 0.1535 0.0589 0.1235
#> $ conf.high: num 0.846 0.957 0.941 0.877
#> $ group : Factor w/ 2 levels "0","1": 1 2 1 2
#> - attr(*, "legend.labels")= chr [1:2] "0" "1"
#> - attr(*, "x.is.factor")= chr "1"
#> - attr(*, "continuous.group")= logi FALSE
#> - attr(*, "rawdata")='data.frame': 12 obs. of 3 variables:
#> ..$ response: Factor w/ 2 levels "0","1": 1 2 2 1 2 1 1 2 1 2 ...
#> ..$ x : num [1:12] 0 0 1 1 0 1 0 1 1 0 ...
#> .. ..- attr(*, "labels")= Named num [1:2] 0 1
#> .. .. ..- attr(*, "names")= chr [1:2] "0" "1"
#> ..$ group : Factor w/ 2 levels "0","1": 2 2 2 1 1 2 1 1 2 2 ...
#> - attr(*, "title")= chr "Predicted probabilities of Choice"
#> - attr(*, "x.title")= chr "LOC"
#> - attr(*, "y.title")= chr "Choice"
#> - attr(*, "legend.title")= chr "SIZE"
#> - attr(*, "x.axis.labels")= chr [1:2] "0" "1"
#> - attr(*, "constant.values")= Named list()
#> - attr(*, "terms")= chr [1:2] "LOC" "SIZE"
#> - attr(*, "original.terms")= chr [1:2] "LOC" "SIZE"
#> - attr(*, "at.list")=List of 2
#> ..$ LOC : chr [1:2] "0" "1"
#> ..$ SIZE: chr [1:2] "0" "1"
#> - attr(*, "ci.lvl")= num 0.95
#> - attr(*, "family")= chr "binomial"
#> - attr(*, "link")= chr "logit"
#> - attr(*, "logistic")= chr "1"
#> - attr(*, "is.trial")= chr "0"
#> - attr(*, "fitfun")= chr "glm"
#> - attr(*, "model.name")= chr "glm.model"
str(glm.model)
#> List of 30
#> $ coefficients : Named num [1:4] -0.693 0.693 1.386 -1.386
#> ..- attr(*, "names")= chr [1:4] "(Intercept)" "LOC1" "SIZE1" "LOC1:SIZE1"
#> $ residuals : Named num [1:12] -3 1.5 2 -2 3 ...
#> ..- attr(*, "names")= chr [1:12] "1" "2" "3" "4" ...
#> $ fitted.values : Named num [1:12] 0.667 0.667 0.5 0.5 0.333 ...
#> ..- attr(*, "names")= chr [1:12] "1" "2" "3" "4" ...
#> $ effects : Named num [1:12] 0.00 2.22e-16 -5.66e-01 -5.66e-01 2.12 ...
#> ..- attr(*, "names")= chr [1:12] "(Intercept)" "LOC1" "SIZE1" "LOC1:SIZE1" ...
#> $ R : num [1:4, 1:4] -1.683 0 0 0 -0.891 ...
#> ..- attr(*, "dimnames")=List of 2
#> .. ..$ : chr [1:4] "(Intercept)" "LOC1" "SIZE1" "LOC1:SIZE1"
#> .. ..$ : chr [1:4] "(Intercept)" "LOC1" "SIZE1" "LOC1:SIZE1"
#> $ rank : int 4
#> $ qr :List of 5
#> ..$ qr : num [1:12, 1:4] -1.683 0.28 0.297 0.297 0.28 ...
#> .. ..- attr(*, "dimnames")=List of 2
#> .. .. ..$ : chr [1:12] "1" "2" "3" "4" ...
#> .. .. ..$ : chr [1:4] "(Intercept)" "LOC1" "SIZE1" "LOC1:SIZE1"
#> ..$ rank : int 4
#> ..$ qraux: num [1:4] 1.28 1.23 1.2 1.33
#> ..$ pivot: int [1:4] 1 2 3 4
#> ..$ tol : num 1e-11
#> ..- attr(*, "class")= chr "qr"
#> $ family :List of 12
#> ..$ family : chr "binomial"
#> ..$ link : chr "logit"
#> ..$ linkfun :function (mu)
#> ..$ linkinv :function (eta)
#> ..$ variance :function (mu)
#> ..$ dev.resids:function (y, mu, wt)
#> ..$ aic :function (y, n, mu, wt, dev)
#> ..$ mu.eta :function (eta)
#> ..$ initialize: language { if (NCOL(y) == 1) { ...
#> ..$ validmu :function (mu)
#> ..$ valideta :function (eta)
#> ..$ simulate :function (object, nsim)
#> ..- attr(*, "class")= chr "family"
#> $ linear.predictors: Named num [1:12] 6.93e-01 6.93e-01 6.66e-16 3.33e-16 -6.93e-01 ...
#> ..- attr(*, "names")= chr [1:12] "1" "2" "3" "4" ...
#> $ deviance : num 16
#> $ aic : num 24
#> $ null.deviance : num 16.6
#> $ iter : int 4
#> $ weights : Named num [1:12] 0.222 0.222 0.25 0.25 0.222 ...
#> ..- attr(*, "names")= chr [1:12] "1" "2" "3" "4" ...
#> $ prior.weights : Named num [1:12] 1 1 1 1 1 1 1 1 1 1 ...
#> ..- attr(*, "names")= chr [1:12] "1" "2" "3" "4" ...
#> $ df.residual : int 8
#> $ df.null : int 11
#> $ y : Named num [1:12] 0 1 1 0 1 0 0 1 0 1 ...
#> ..- attr(*, "names")= chr [1:12] "1" "2" "3" "4" ...
#> $ converged : logi TRUE
#> $ boundary : logi FALSE
#> $ model :'data.frame': 12 obs. of 3 variables:
#> ..$ Choice: Factor w/ 2 levels "0","1": 1 2 2 1 2 1 1 2 1 2 ...
#> ..$ LOC : Factor w/ 2 levels "0","1": 1 1 2 2 1 2 1 2 2 1 ...
#> ..$ SIZE : Factor w/ 2 levels "0","1": 2 2 2 1 1 2 1 1 2 2 ...
#> ..- attr(*, "terms")=Classes 'terms', 'formula' language Choice ~ LOC * SIZE
#> .. .. ..- attr(*, "variables")= language list(Choice, LOC, SIZE)
#> .. .. ..- attr(*, "factors")= int [1:3, 1:3] 0 1 0 0 0 1 0 1 1
#> .. .. .. ..- attr(*, "dimnames")=List of 2
#> .. .. .. .. ..$ : chr [1:3] "Choice" "LOC" "SIZE"
#> .. .. .. .. ..$ : chr [1:3] "LOC" "SIZE" "LOC:SIZE"
#> .. .. ..- attr(*, "term.labels")= chr [1:3] "LOC" "SIZE" "LOC:SIZE"
#> .. .. ..- attr(*, "order")= int [1:3] 1 1 2
#> .. .. ..- attr(*, "intercept")= int 1
#> .. .. ..- attr(*, "response")= int 1
#> .. .. ..- attr(*, ".Environment")=<environment: R_GlobalEnv>
#> .. .. ..- attr(*, "predvars")= language list(Choice, LOC, SIZE)
#> .. .. ..- attr(*, "dataClasses")= Named chr [1:3] "factor" "factor" "factor"
#> .. .. .. ..- attr(*, "names")= chr [1:3] "Choice" "LOC" "SIZE"
#> $ call : language glm(formula = Choice ~ LOC * SIZE, family = binomial(link = "logit"), data = cedata.1)
#> $ formula :Class 'formula' language Choice ~ LOC * SIZE
#> .. ..- attr(*, ".Environment")=<environment: R_GlobalEnv>
#> $ terms :Classes 'terms', 'formula' language Choice ~ LOC * SIZE
#> .. ..- attr(*, "variables")= language list(Choice, LOC, SIZE)
#> .. ..- attr(*, "factors")= int [1:3, 1:3] 0 1 0 0 0 1 0 1 1
#> .. .. ..- attr(*, "dimnames")=List of 2
#> .. .. .. ..$ : chr [1:3] "Choice" "LOC" "SIZE"
#> .. .. .. ..$ : chr [1:3] "LOC" "SIZE" "LOC:SIZE"
#> .. ..- attr(*, "term.labels")= chr [1:3] "LOC" "SIZE" "LOC:SIZE"
#> .. ..- attr(*, "order")= int [1:3] 1 1 2
#> .. ..- attr(*, "intercept")= int 1
#> .. ..- attr(*, "response")= int 1
#> .. ..- attr(*, ".Environment")=<environment: R_GlobalEnv>
#> .. ..- attr(*, "predvars")= language list(Choice, LOC, SIZE)
#> .. ..- attr(*, "dataClasses")= Named chr [1:3] "factor" "factor" "factor"
#> .. .. ..- attr(*, "names")= chr [1:3] "Choice" "LOC" "SIZE"
#> $ data :'data.frame': 12 obs. of 7 variables:
#> ..$ id : num [1:12] 1 1 1 1 1 1 2 2 2 2 ...
#> ..$ QES : num [1:12] 1 1 2 2 3 3 1 1 2 2 ...
#> ..$ Alt : num [1:12] 1 2 1 2 1 2 1 2 1 2 ...
#> ..$ LOC : Factor w/ 2 levels "0","1": 1 1 2 2 1 2 1 2 2 1 ...
#> ..$ SIZE : Factor w/ 2 levels "0","1": 2 2 2 1 1 2 1 1 2 2 ...
#> ..$ Choice: Factor w/ 2 levels "0","1": 1 2 2 1 2 1 1 2 1 2 ...
#> ..$ gender: num [1:12] 1 1 1 1 1 1 0 0 0 0 ...
#> $ offset : NULL
#> $ control :List of 3
#> ..$ epsilon: num 1e-08
#> ..$ maxit : num 25
#> ..$ trace : logi FALSE
#> $ method : chr "glm.fit"
#> $ contrasts :List of 2
#> ..$ LOC : chr "contr.treatment"
#> ..$ SIZE: chr "contr.treatment"
#> $ xlevels :List of 2
#> ..$ LOC : chr [1:2] "0" "1"
#> ..$ SIZE: chr [1:2] "0" "1"
#> - attr(*, "class")= chr [1:2] "glm" "lm"
Created on 2020-09-13 by the reprex package (v0.3.0)
The two objects do provide all the pieces required to produce data frames for rendering as kable or pander tables.