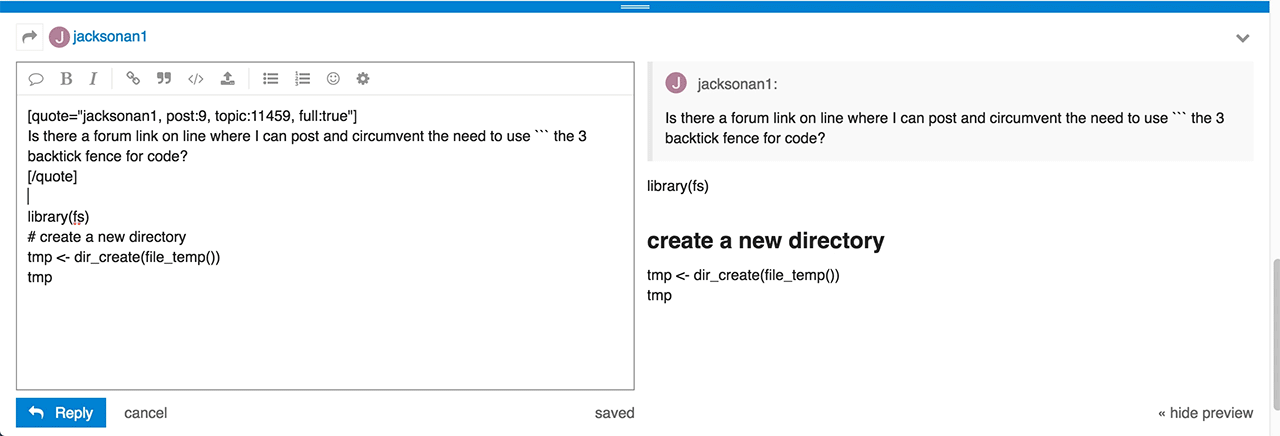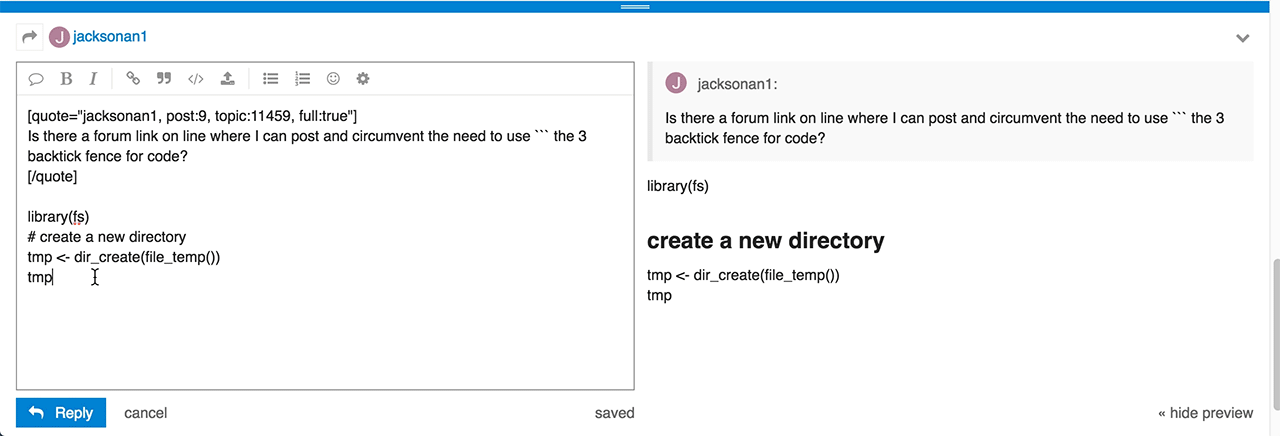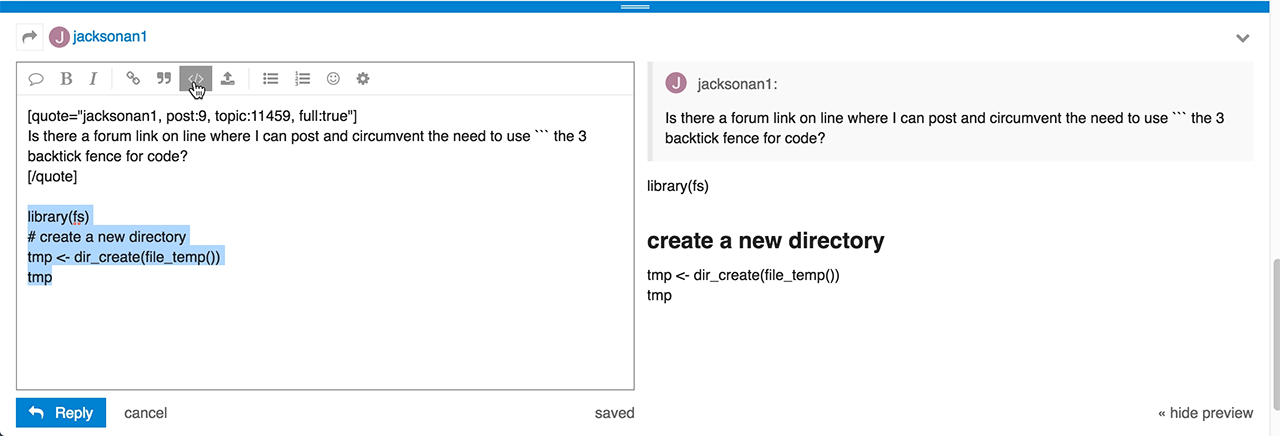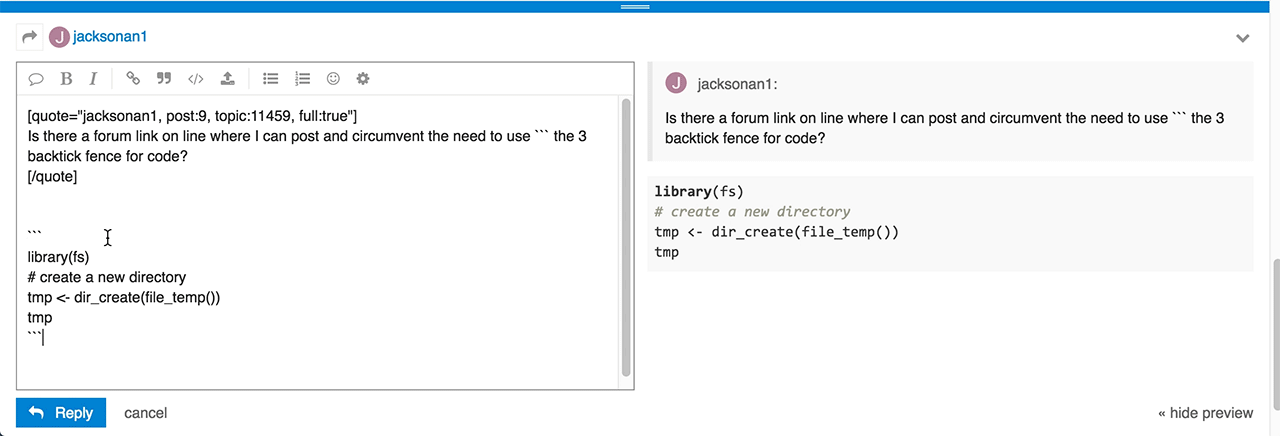
Do you have any idea of what I did wrong and how to correct?
I'm still at a bit of a loss to understand what you are asking.
In an earlier post you asked:
My question is do I have to put in the entire table as you did or can I just designate the junk entry?
read.table reads from a file or, in my example case, a text string, into a data.frame. I don't understand what you mean by this question. My code showed how to skip the first line which was not required, and then read the file. Which was what you seemed to be trying to do.
Then you tried this:
datr<-read.table(file="popfo7n_100.fit",skip=1, nrows=46400, header=T)
which, if I understand you correctly, has done what you wanted except that the character variables have been stored as factors. Yes?
I presume you have more than 46400 rows of data but only want to read 46400 rows. If you are reading the complete file, the nrows parameter is redundant. As an aside, if you do have more than 46400 rows, it may be better to read the whole file and then subset the data.frame based on whatever defines those first 46400 rows as being of interest (more robust if revisited in the future - you may not need to know the number 46400).
I do not understand why, having got a read.table command that works, you then seem to be trying to read the same file without the skip=1. This will cause the line of garbage to be read and interpreted as headers which has implications for the number of columns read.table will expect to find in the following lines.
Try:
datr<-read.table(file = "popfo7n_100.fit", header = TRUE,
skip = 1, stringsAsFactors = FALSE)
with or without the "nrows = 46400", as appropriate.
Without access to your file; so not knowing what the first line actually says (number of words), or how many columns of data there are, or how they're are separated, I'm not sure why you only got 1 row with your last attempt. You got more than 1 row when using skip=1 and specifying nrows?
It could possibly be to do with the way you've called read.table or it may be related to the origin of the data file. Is it a text file that was created on Windows that you are attempting to read on a Mac, or vice versa?