Cross-posted at
I am having some trouble with an error message I can't solve.
I use the following code in to run a Kruskal Wallis in R and then do a pairwise comparison using Dunn's:
res.kruskal <- mydata %>% kruskal_test(values ~ group)
res.kruskal
stat.test <- mydata %>% dunn_test(values ~ group, p.adjust.method = "hochberg")
stat.test
stat.test <- stat.test %>% add_xy_position(x = "group")
I then use this result to plot out the significances like so:
ggboxplot(mydata, x = "group", y = "values, fill = "group") +
stat_pvalue_manual(stat.test, hide.ns = FALSE)
And get is nice looking plot (hooray!)
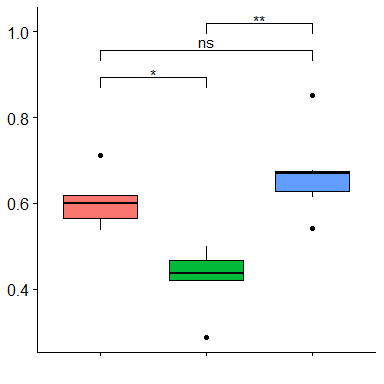
But the Cover Test is apparently more powerful and preferred over Dunn's... (any opinions on this would be welcome too!). Running the following code returns an error:
res.kruskal <- immdatamed %>% kruskal_test(LplastinTL ~ group)
res.kruskal
attach(immdatamed)
stat.test <- conover.test(LplastinTL, group, method = "hochberg")
stat.test
stat.test <- stat.test %>% add_xy_position(x = "group")
detach(immdatamed)
stat.test <- stat.test %>% add_xy_position(x = "group")
Error in asserttat_group_columns_exists(test) :
data should contain group1 and group2 columns
I can't figure this out... I can run the Conover test fine but can't solve the above error to get it to work with ggboxplot.
I would actually prefer to use the more aesthetically pleasing ggplot & geom_boxplot but can't get that to interface with Dunn_test at all...
Any solutions would be most welcome!
Thank you
P.S. I was using this with ggplot: Add P-values and Significance Levels to ggplots - Articles - STHDA but I cont want a wilcoxon or t.test for pairwise comparison...