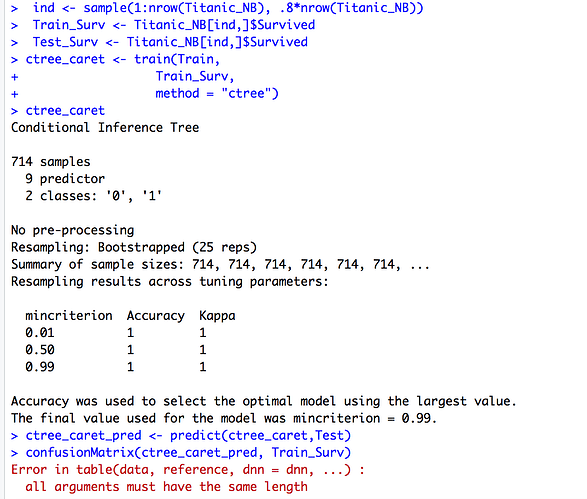
Hey guys, sorry to bother you all but I am having trouble with a school project, I am trying to create a naives bayes model and decision tree for a database. I keep getting an error and for some reason when I do my prediction for both ctree and naive bayes, I get different scores of accuracies? After that i cannot seem to get the confusion matrix for both sets?
Please and thanks!
Hi, and welcome!
Please see the FAQ: What's a reproducible example (`reprex`) and how do I create one? Using a reprex, complete with representative data will attract quicker and more answers. And also the homework policy.
Screenshots are not very helpful because it takes a fair amount of effort to reverse engineer the problem. With a reprex it's just cut and paste, if the data is included. In this case, there's the impediment of finding Titanic_NB. We don't know if it's the same as the standard Titanic dataset.
Without the reprex, all to be done is to look at the function signature for confusionMarix to see what it expects of its argument, and try an example from the documentation.
One of the hard things to get used to in R is the concept that everything is an object that has properties. Some objects have properties that allow them to operate on other objects to produce new objects. Those are functions.
Think of R as school algebra writ large: f(x) = y, where the objects are f, a function, x, an object (and there may be several) termed the argument and y is an object termed a value, which can be as simple as a single number (aka an atomic vector) or a very packed object with a multitude of data and labels.
And, because functions are also objects, they can be arguments to other functions, like the old g(f(x)) = y. (Trivia, this is called being a first class object.)
Although there are function objects in R that operate like control statements in imperative/procedural language, they are best used "under the hood." As it presents to users interactively, R is a functional programming language. Instead of saying
take this, take that, do this, then do that, then if the result is this one thing, do this other thing, but if not do something else and give me the answer
in the style of most common programming languages. But R allows the user to say simply
use this function to take this argument and turn it into the value I want for a result
That's powerful!
And it's also the key to unpacking the notorious mysterious help pages.
So, let's skim help(confusionMatrix)
The signature is
confusionMatrix(data, ...)
The first argument (sometimes called a parameter by analogy to other areas) is, well, data with the second, the mysterious \ldots.
Quick aside, data is the name of a built in object, and if you use it you risk what's called namespace collision or at least confusion, like a host of an age cohort all with the same given name. To check if your preferred object name is already taken, just
data
#> function (..., list = character(), package = NULL, lib.loc = NULL,
#> verbose = getOption("verbose"), envir = .GlobalEnv, overwrite = TRUE)
# HUGE SNIP HERE
#> <environment: namespace:utils>
my_data
#> Error in eval(expr, envir, enclos): object 'my_data' not found
Created on 2020-04-04 by the reprex package (v0.3.0)
Easy to see which one you want.
Ok, so what should data be?
data \ \ \ a factor of predicted classes (for the default method) or an object of class table.
That tell us right there that whatever we feed as the first argument has to be either a factor or a data table.
And what of \ldots?
\ldots\ \ \ options to be passed to table. NOTE: do not include dnn here
So, we only get to use \ldots if we are feeding confusionMatrix a data table object.
To find out about an object, there's the str() structure command. The bests way to understand it is to work an example from the help(confusionMatrix) page and pause along the way to insert it.
library(caret)
#> Loading required package: lattice
#> Loading required package: ggplot2
lvs <- c("normal", "abnormal")
str(lvl)
#> Error in str(lvl): object 'lvl' not found
truth <- factor(rep(lvs, times = c(86, 258)),
levels = rev(lvs))
str(truth)
#> Factor w/ 2 levels "abnormal","normal": 2 2 2 2 2 2 2 2 2 2 ...
pred <- factor(
c(
rep(lvs, times = c(54, 32)),
rep(lvs, times = c(27, 231))),
levels = rev(lvs))
str(pred)
#> Factor w/ 2 levels "abnormal","normal": 2 2 2 2 2 2 2 2 2 2 ...
xtab <- table(pred, truth)
str(xtab)
#> 'table' int [1:2, 1:2] 231 27 32 54
#> - attr(*, "dimnames")=List of 2
#> ..$ pred : chr [1:2] "abnormal" "normal"
#> ..$ truth: chr [1:2] "abnormal" "normal"
a <- confusionMatrix(xtab)
str(a)
#> List of 6
#> $ positive: chr "abnormal"
#> $ table : 'table' int [1:2, 1:2] 231 27 32 54
#> ..- attr(*, "dimnames")=List of 2
#> .. ..$ pred : chr [1:2] "abnormal" "normal"
#> .. ..$ truth: chr [1:2] "abnormal" "normal"
#> $ overall : Named num [1:7] 0.828 0.534 0.784 0.867 0.75 ...
#> ..- attr(*, "names")= chr [1:7] "Accuracy" "Kappa" "AccuracyLower" "AccuracyUpper" ...
#> $ byClass : Named num [1:11] 0.895 0.628 0.878 0.667 0.878 ...
#> ..- attr(*, "names")= chr [1:11] "Sensitivity" "Specificity" "Pos Pred Value" "Neg Pred Value" ...
#> $ mode : chr "sens_spec"
#> $ dots : list()
#> - attr(*, "class")= chr "confusionMatrix"
b <- confusionMatrix(pred, truth)
str(b)
#> List of 6
#> $ positive: chr "abnormal"
#> $ table : 'table' int [1:2, 1:2] 231 27 32 54
#> ..- attr(*, "dimnames")=List of 2
#> .. ..$ Prediction: chr [1:2] "abnormal" "normal"
#> .. ..$ Reference : chr [1:2] "abnormal" "normal"
#> $ overall : Named num [1:7] 0.828 0.534 0.784 0.867 0.75 ...
#> ..- attr(*, "names")= chr [1:7] "Accuracy" "Kappa" "AccuracyLower" "AccuracyUpper" ...
#> $ byClass : Named num [1:11] 0.895 0.628 0.878 0.667 0.878 ...
#> ..- attr(*, "names")= chr [1:11] "Sensitivity" "Specificity" "Pos Pred Value" "Neg Pred Value" ...
#> $ mode : chr "sens_spec"
#> $ dots : list()
#> - attr(*, "class")= chr "confusionMatrix"
c <- confusionMatrix(xtab, prevalence = 0.25)
str(c)
#> List of 6
#> $ positive: chr "abnormal"
#> $ table : 'table' int [1:2, 1:2] 231 27 32 54
#> ..- attr(*, "dimnames")=List of 2
#> .. ..$ pred : chr [1:2] "abnormal" "normal"
#> .. ..$ truth: chr [1:2] "abnormal" "normal"
#> $ overall : Named num [1:7] 0.828 0.534 0.784 0.867 0.75 ...
#> ..- attr(*, "names")= chr [1:7] "Accuracy" "Kappa" "AccuracyLower" "AccuracyUpper" ...
#> $ byClass : Named num [1:11] 0.895 0.628 0.445 0.947 0.878 ...
#> ..- attr(*, "names")= chr [1:11] "Sensitivity" "Specificity" "Pos Pred Value" "Neg Pred Value" ...
#> $ mode : chr "sens_spec"
#> $ dots : list()
#> - attr(*, "class")= chr "confusionMatrix"
Created on 2020-04-04 by the reprex package (v0.3.0)
Try doing that to your code and see what you find.
This topic was automatically closed 21 days after the last reply. New replies are no longer allowed.