I’m using mlogit to estimate a regression model. Further, i’m using sjPlot to assess marginal effects and predicted probabilities from said model (according to package version 2.6.3 "sjPlot" explicitly supports "mlogit")
I’ve run into a problem plotting/assessing MargEff/PredProb (via sjPlot). I keep getting this error:
plot_model(ce.model, type = "pred", terms = "SIZE")
Error in contrasts<-(tmp, value = contr.funs[1 + isOF[nn]]) : ***
*** contrasts can be applied only to factors with 2 or more levels
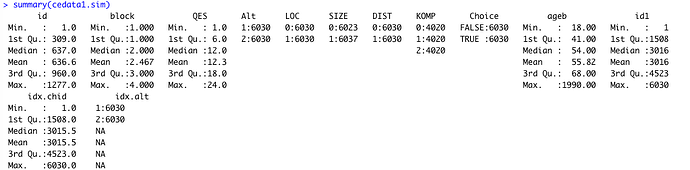
Review of several forum posts (see below) suggests that this is likely related to NAs in dataset but, although there are “NAs” in the relevant variables in my regression model, there are in fact NAs in the indexing columns that “mlogit.data” creates in order to run mlogit (see attached screen shot, and other details pasted below).
So the question is: Assuming these indexed NAs are causing my error message, how can I remove them but still rely on mlogit for my regression? Are there other work arounds?
(since my question is conceptual I have not included a reprex, but i could if needed…)
Scott
MY REVIEW OF FORUM POSTS ...
... led me to troubleshoot all of the following:
- factor levels of an explanatory variables are not differentiated, i.e., only 1 level exists (including possible “empty” levels).
- NAs exists somewhere in dataset which knocks out a bunch of factor levels from dataset
- Existence of exotic characters å, ä, ä in dataset.
- other variables not used in regression, but included in dataset that may cause problems…
(see here database - Error in contrasts when defining a linear model in R - Stack Overflow )
DETAILS ON MLOGIT.data
To use mlogit, one must first use mlogit.data to convert the dataframe to something that mlogit undertands. In short, it means defining index variables (e.g., which variable is the “choice” variable? How many alternatives did each individual face in the panel data? etc).
see attached screenshot. the “idx.alt” column has length=2 and a bunch of NAs. The last 3 columns are all indexes (id1, idx.chid, idx.alt) from “mlogit.data” command
Several posts related to "mlogit.data", but none seem to encounter my specific problem.
See also info mlogit.data here: https://cran.r-project.org/web/packages/mlogit/vignettes/c2.formula.data.html