The Gini Coefficient
The Gini Coefficient is a measure of equality that ranges from 0 (perfect equality) to 1 (a single entity has 100% of some quantity). In a closed post, I suggested that because Gini was a continuous variable, capable of taking on any value between 0 and 1, ordinary least square linear regression would be a good place to start. (Although if it were binary 0 or 1, logistic regression using the binomial family would be required.)
A sidebar
@Yarnabrina and I had a message discussion whether a Gini observation of exactly 0 or exactly 1 would generate a correlation of -Inf or +Inf and, more generally, whether it is guaranteed that predicted results will be bounded by the interval of the dependent variable.
I made the argument from practicality -- the low probability of occurrence justifies starting with the assumption that the data don't contain values of exactly 0 or 1.
A toy model
dat <- runif(1000000)
x <- sample(dat, 1000)
y <- sample(dat, 1000) * sample(dat,1000) * 10
head(x)
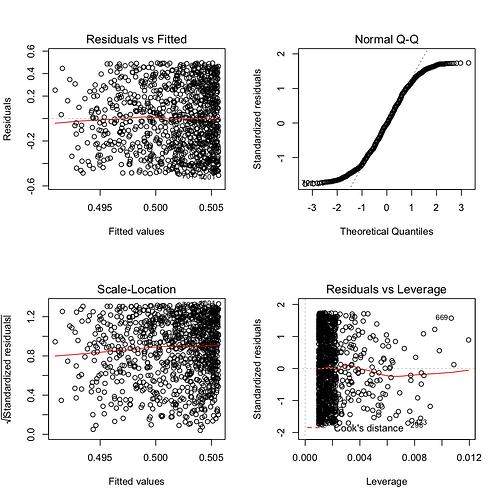
mod <- lm(x ~ y)
summary(mod)
##
## Call:
## lm(formula = x ~ y)
##
## Residuals:
## Min 1Q Median 3Q Max
## -0.50109 -0.23728 -0.00731 0.23897 0.51709
##
## Coefficients:
## Estimate Std. Error t value Pr(>|t|)
## (Intercept) 0.477383 0.013655 34.959 <2e-16 ***
## y 0.004134 0.004059 1.018 0.309
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Residual standard error: 0.2856 on 998 degrees of freedom
## Multiple R-squared: 0.001038, Adjusted R-squared: 3.698e-05
## F-statistic: 1.037 on 1 and 998 DF, p-value: 0.3088
This is exactly what we would expect to see in regressing a random dependent variable on a random independent variable almost all the time. The intercept is near 0.5 and the coefficient near zero.
A toy pathological case
Now let's see if introducing 0 and 1 into x & y produces any different result
dat <- runif(1000000)
x <- sample(dat, 1000)
x <- c(x,0,1)
y <- sample(dat, 1000) * sample(dat,1000) * 10
y <- c(y,0,1)
mod <- lm(x ~ y)
summary(mod)
##
## Call:
## lm(formula = x ~ y)
##
## Residuals:
## Min 1Q Median 3Q Max
## -0.50373 -0.25385 -0.00786 0.26139 0.51217
##
## Coefficients:
## Estimate Std. Error t value Pr(>|t|)
## (Intercept) 0.484740 0.013746 35.265 <2e-16 ***
## y 0.003093 0.004028 0.768 0.443
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Residual standard error: 0.2893 on 1000 degrees of freedom
## Multiple R-squared: 0.0005895, Adjusted R-squared: -0.0004099
## F-statistic: 0.5898 on 1 and 1000 DF, p-value: 0.4427
No substantive difference is visible.
Are OLS predictions theoretically always within the bounds of the dependant variable?
Peter Dalgaard, Introductory Statistics with R
The linear regression model is provided by
y_i = \alpha + \beta x_i + \epsilon_i
in which the \epsilon_i are assumed independent an N(0,\sigma^2). The nonrandom part of the equation describes the y_i as lying on a straight line. The slope of the line (the regression coefficient) is \beta, the increase per unit change in x. The line intersects the y axis at the intercept \alpha.
at 109.
He then goes on to explain that the method of least squares can be used to estimate \alpha, \beta, \sigma^2 by choosing \alpha, \beta to minimize the sum of squared residuals.
Even if max(x) is paired with max(y) in the data and y \gg x, the residual is still measured relative to the slope, which terminates at max(x) because \ni x > max(x).
I don't see any way of falsifying that conclusion, but my formal training is limited and I invite criticism.