I hope I grabbed the correct parts of the code. The following works for me. Try copying it and see if it works for you.
summarySE <- function(data=NULL, measurevar, groupvars=NULL, na.rm=FALSE,
conf.interval=.95, .drop=TRUE) {
library(plyr)
# New version of length which can handle NA's: if na.rm==T, don't count them
length2 <- function (x, na.rm=FALSE) {
if (na.rm) sum(!is.na(x))
else length(x)
}
# This does the summary. For each group's data frame, return a vector with
# N, mean, and sd
datac <- ddply(data, groupvars, .drop=.drop,
.fun = function(xx, col) {
c(N = length2(xx[[col]], na.rm=na.rm),
mean = mean (xx[[col]], na.rm=na.rm),
sd = sd (xx[[col]], na.rm=na.rm)
)
},
measurevar
)
# Rename the "mean" column
datac <- rename(datac, c("mean" = measurevar))
datac$se <- datac$sd / sqrt(datac$N) # Calculate standard error of the mean
# Confidence interval multiplier for standard error
# Calculate t-statistic for confidence interval:
# e.g., if conf.interval is .95, use .975 (above/below), and use df=N-1
ciMult <- qt(conf.interval/2 + .5, datac$N-1)
datac$ci <- datac$se * ciMult
return(datac)
}
################
normDataWithin <- function(data=NULL, idvar, measurevar, betweenvars=NULL,
na.rm=FALSE, .drop=TRUE) {
library(plyr)
# Measure var on left, idvar + between vars on right of formula.
data.subjMean <- ddply(data, c(idvar, betweenvars), .drop=.drop,
.fun = function(xx, col, na.rm) {
c(subjMean = mean(xx[,col], na.rm=na.rm))
},
measurevar,
na.rm
)
# Put the subject means with original data
data <- merge(data, data.subjMean)
# Get the normalized data in a new column
measureNormedVar <- paste(measurevar, "_norm", sep="")
data[,measureNormedVar] <- data[,measurevar] - data[,"subjMean"] +
mean(data[,measurevar], na.rm=na.rm)
# Remove this subject mean column
data$subjMean <- NULL
return(data)
}
################
summarySEwithin <- function(data=NULL, measurevar, betweenvars=NULL, withinvars=NULL,
idvar=NULL, na.rm=FALSE, conf.interval=.95, .drop=TRUE) {
# Ensure that the betweenvars and withinvars are factors
factorvars <- vapply(data[, c(betweenvars, withinvars), drop=FALSE],
FUN=is.factor, FUN.VALUE=logical(1))
if (!all(factorvars)) {
nonfactorvars <- names(factorvars)[!factorvars]
message("Automatically converting the following non-factors to factors: ",
paste(nonfactorvars, collapse = ", "))
data[nonfactorvars] <- lapply(data[nonfactorvars], factor)
}
# Get the means from the un-normed data
datac <- summarySE(data, measurevar, groupvars=c(betweenvars, withinvars),
na.rm=na.rm, conf.interval=conf.interval, .drop=.drop)
# Drop all the unused columns (these will be calculated with normed data)
datac$sd <- NULL
datac$se <- NULL
datac$ci <- NULL
# Norm each subject's data
ndata <- normDataWithin(data, idvar, measurevar, betweenvars, na.rm, .drop=.drop)
# This is the name of the new column
measurevar_n <- paste(measurevar, "_norm", sep="")
# Collapse the normed data - now we can treat between and within vars the same
ndatac <- summarySE(ndata, measurevar_n, groupvars=c(betweenvars, withinvars),
na.rm=na.rm, conf.interval=conf.interval, .drop=.drop)
# Apply correction from Morey (2008) to the standard error and confidence interval
# Get the product of the number of conditions of within-S variables
nWithinGroups <- prod(vapply(ndatac[,withinvars, drop=FALSE], FUN=nlevels,
FUN.VALUE=numeric(1)))
correctionFactor <- sqrt( nWithinGroups / (nWithinGroups-1) )
# Apply the correction factor
ndatac$sd <- ndatac$sd * correctionFactor
ndatac$se <- ndatac$se * correctionFactor
ndatac$ci <- ndatac$ci * correctionFactor
# Combine the un-normed means with the normed results
merge(datac, ndatac)
}
##############
data <- read.table(header=TRUE, text='
Subject RoundMono SquareMono RoundColor SquareColor
1 41 40 41 37
2 57 56 56 53
3 52 53 53 50
4 49 47 47 47
5 47 48 48 47
6 37 34 35 36
7 47 50 47 46
8 41 40 38 40
9 48 47 49 45
10 37 35 36 35
11 32 31 31 33
12 47 42 42 42
')
# Convert it to long format
library(reshape2)
data_long <- melt(data=data, id.var="Subject",
measure.vars=c("RoundMono", "SquareMono", "RoundColor", "SquareColor"),
variable.name="Condition")
names(data_long)[names(data_long)=="value"] <- "Time"
# Split Condition column into Shape and ColorScheme
data_long$Shape <- NA
data_long$Shape[grepl("^Round", data_long$Condition)] <- "Round"
data_long$Shape[grepl("^Square", data_long$Condition)] <- "Square"
data_long$Shape <- factor(data_long$Shape)
data_long$ColorScheme <- NA
data_long$ColorScheme[grepl("Mono$", data_long$Condition)] <- "Monochromatic"
data_long$ColorScheme[grepl("Color$", data_long$Condition)] <- "Colored"
data_long$ColorScheme <- factor(data_long$ColorScheme, levels=c("Monochromatic","Colored"))
# Remove the Condition column now
data_long$Condition <- NULL
# Look at first few rows
head(data_long)
#> Subject Time Shape ColorScheme
#> 1 1 41 Round Monochromatic
#> 2 2 57 Round Monochromatic
#> 3 3 52 Round Monochromatic
#> 4 4 49 Round Monochromatic
#> 5 5 47 Round Monochromatic
#> 6 6 37 Round Monochromatic
datac <- summarySEwithin(data_long, measurevar="Time", withinvars=c("Shape","ColorScheme"), idvar="Subject")
datac
#> Shape ColorScheme N Time Time_norm sd se ci
#> 1 Round Colored 12 43.58333 43.58333 1.212311 0.3499639 0.7702654
#> 2 Round Monochromatic 12 44.58333 44.58333 1.331438 0.3843531 0.8459554
#> 3 Square Colored 12 42.58333 42.58333 1.461630 0.4219364 0.9286757
#> 4 Square Monochromatic 12 43.58333 43.58333 1.261312 0.3641095 0.8013997
library(ggplot2)
ggplot(datac, aes(x=Shape, y=Time, fill=ColorScheme)) +
geom_bar(position=position_dodge(.9), colour="black", stat="identity") +
geom_errorbar(position=position_dodge(.9), width=.25, aes(ymin=Time-ci, ymax=Time+ci)) +
coord_cartesian(ylim=c(40,46)) +
scale_fill_manual(values=c("#CCCCCC","#FFFFFF")) +
scale_y_continuous(breaks=seq(1:100)) +
theme_bw() +
geom_hline(yintercept=38)
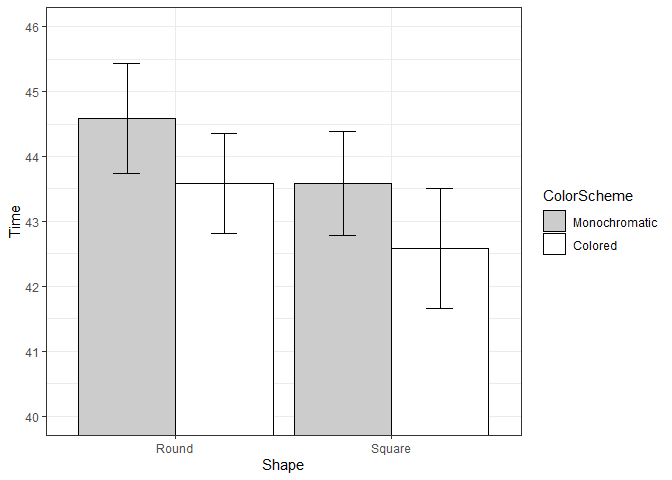
Created on 2020-05-03 by the reprex package (v0.3.0)
