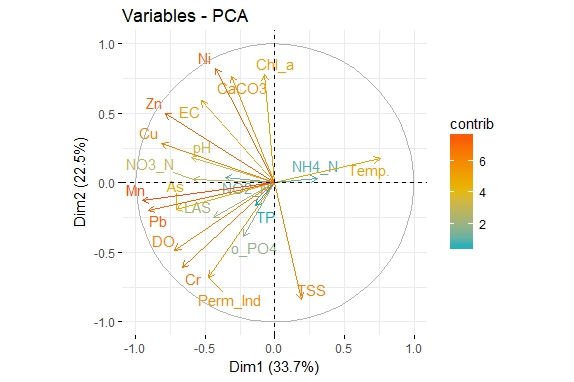
How can I invert/flip of PCA graph, please?
Here is the code:
library("FactoMineR")
library("factoextra")
mydata<- read.csv("Ikizdere.csv", TRUE, ",")
mydata[is.na(mydata)]=0
attach(mydata)
X=cbind (Temp., pH, EC, Perm_Ind, DO, CaCO3, o_PO4, TP, TSS, NO3_N, NO2_N, Chl_a, NH4_N, LAS, As, Cr, Cu, Mn, Ni, Pb, Zn)
summary(X)
cor(X)
res.pca <- princomp(X, scores=TRUE, cor=TRUE)
summary(res.pca)
fviz_pca_var(res.pca, col.var="contrib",
gradient.cols = c("#00AFBB", "#E7B800", "#FC4E07"),
repel = TRUE # Avoid text overlapping
)
jdlong
April 23, 2018, 1:54pm
2
what exactly do you mean by invert/flip? Do you want to sort the y axis in the opposite direction? Reverse the x axis?
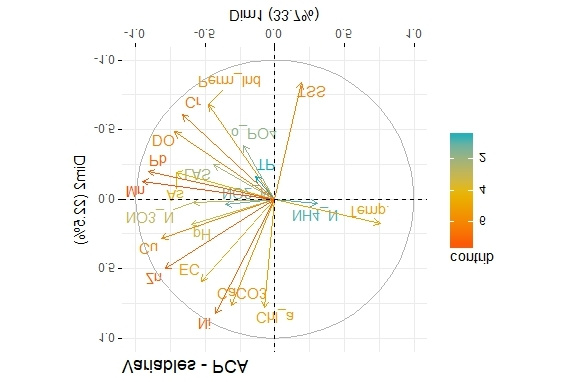
Because what you asked could be interpreted as this:
which is may or may not be what you want.
Thanks for the kind reply. From here you can check my two graphs (https://drive.google.com/file/d/1NzBy5qix7DOCqQ6F88J2aEtga-sp-lY5/view?usp=sharing )
Both graphs are same (of course with same meaning) but the direction of the vectors is in the opposite direction. I want to invert the first plot.
Please download the file so you could see both plots.
jdlong
April 23, 2018, 8:56pm
5
the direction of the vectors has meaning and it hardly seems something one would arbitrarily reverse. But you could swap the axes like this:
fviz_pca_var(res.pca, axes=c(2,1), col.var="contrib",
gradient.cols = c("#00AFBB", "#E7B800", "#FC4E07"),
repel = TRUE # Avoid text overlapping
)
I may be late in answering this, but I noticed the same difference when plot my PCA, wherein the axes were flipped depending on whether I used vegan or factominer.
A quick solution is to add + xlim(c(1,-1) + ylim(c(1,-1)). For instance,
fviz_pca_var(res.PCA, axes=c(1,2), repel=TRUE, col.var="contrib", gradient.cols=c("white","#f47a27 ", "#ed4040 "), ggtheme=theme_minimal()) + xlim(c(1,-1) + ylim(c(1,-1))
2 Likes