Hello,
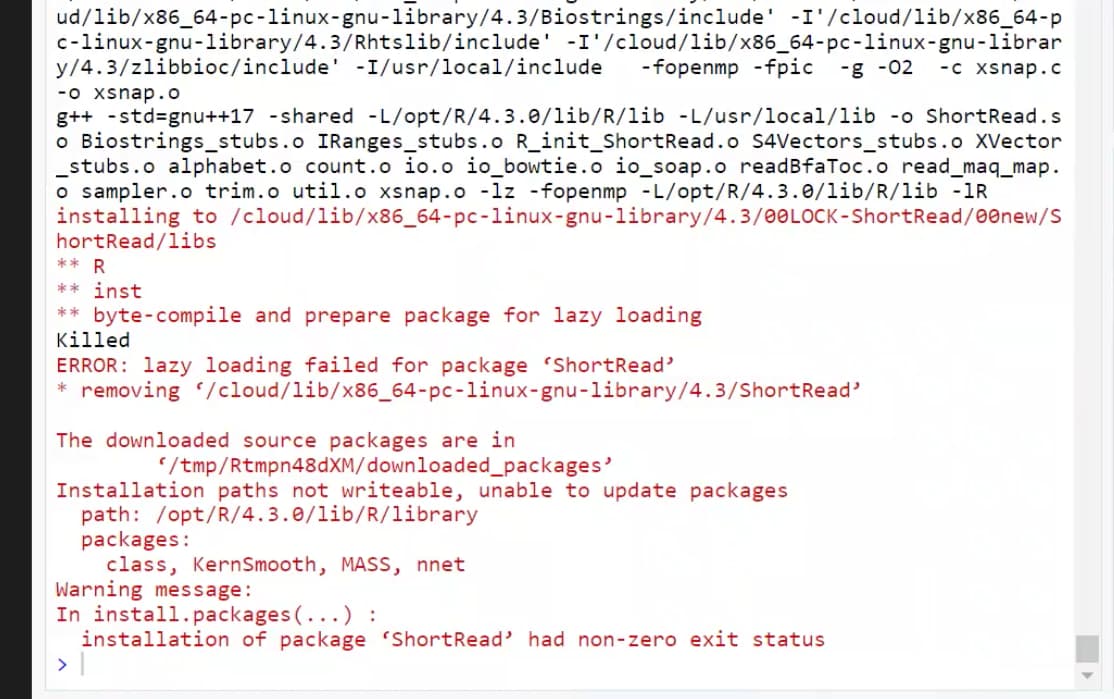
The installation of a Bioconductor package ShortRead in posit.cloud (free tier) fails due to apparent lack of permissions to write to /opt/R/4.3.0/lib/R/library/. I attach a screenshot of the error message (apologies for this, but I am solving someone else's problem via screenshare).
The project is https://posit.cloud/content/5941822 the command to reproduce the error is:
if (!require("BiocManager", quietly = TRUE))
install.packages("BiocManager")
BiocManager::install("ShortRead")
.libPaths() is:
/cloud/lib/x86_64-pc-linux-gnu-library/4.3
/opt/R/4.3.0/lib/R/library/
I should also add that this error happened after a successful installation of CRAN package tidyverse and Bioconductor package Biostrings. Bioconductor package ShortRead is required in addition to Biostrings to install another CRAN package, SimRAD.
Any advice on how to fix this would be very much appreciated!
FJ
Gabor
May 10, 2023, 4:01pm
2
That's probably expected, and you cannot update packages in the system library. So don't update them. Or install their new new versions in your user package library, if you really need to update them.
Thanks. Do I then understand correctly that not much can be done? Unless the package is forced to install in /cloud/lib/x86_64-pc-linux-gnu-library/4.3 rather than /opt/R/4.3.0/lib/R/library/ (which I believe I cannot do)?
Best
Gabor
May 15, 2023, 5:15pm
4
You can probably install these packages to .libPaths()[1]:
install.packages(
c("class", "KernSmooth", "MASS", "nnet"),
lib = .libPaths()[1]
)
But again, these packages receive minimal updates, and it is most likely that you are fine with the versions you already have and you don't actually need to update them.
I am stuck with similar error while deploying shiny io.
2023-05-17T08:20:29.023272+00:00 shinyapps[9119680]: Bioconductor version 3.17 (BiocManager 1.30.20), R 4.3.0 (2023-04-21)force = TRUE to re-install: 'BiocVersion'Bioconductor - 3.17 Software Packages Bioconductor - 3.17 AnnotationData Packages Bioconductor - 3.17 ExperimentData Packages Bioconductor - 3.17 Workflow Packages https://bioconductor.org/packages/3.17/books https://cloud.r-project.org
Gabor
May 17, 2023, 10:31am
6
@smruthi I suggest that you create a new question with the shinyapps.io tag. Does your package call BiocManager::install() explicitly?
1 Like
Thank you very much. Yes, it was called explicitly. Removed it and it works better.
1 Like
system
June 7, 2023, 10:59am
8
This topic was automatically closed 21 days after the last reply. New replies are no longer allowed.