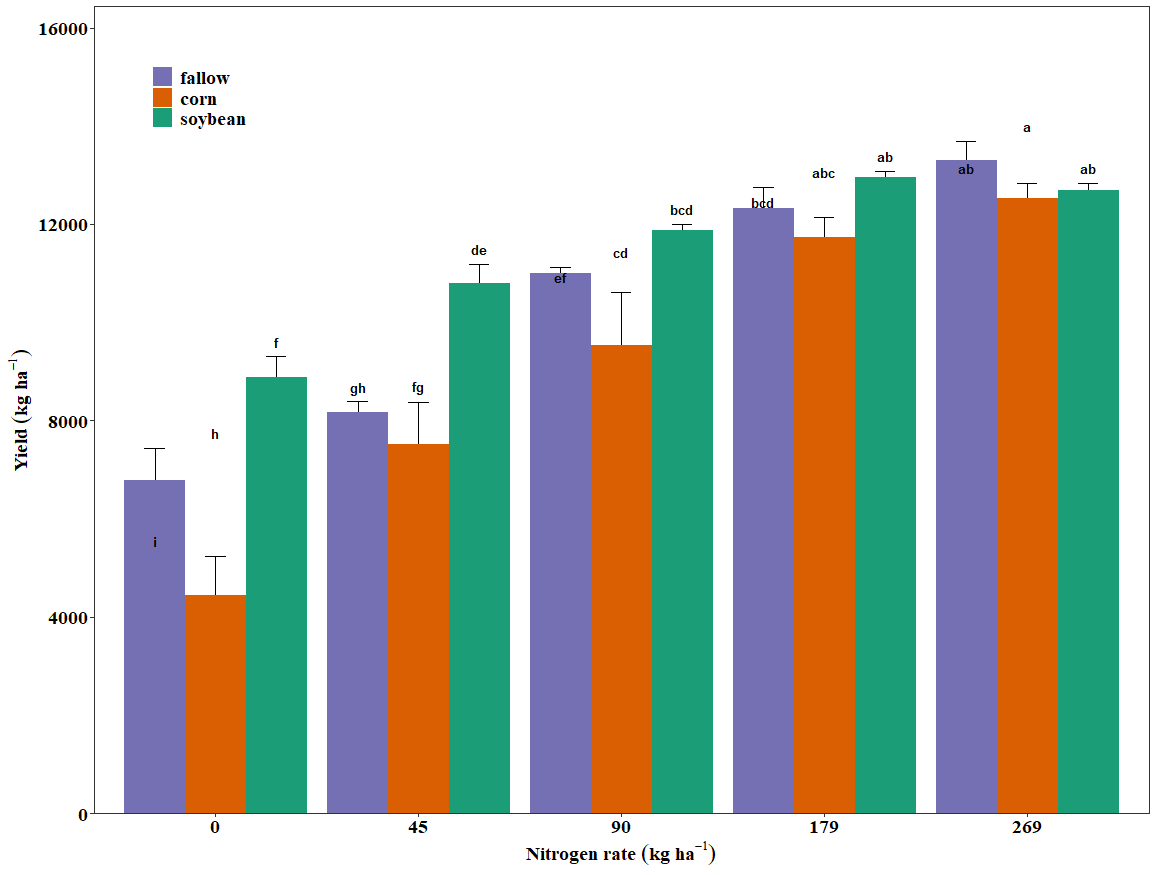
Hi everyone please I can someone help me with this; I need the letters to be at the top of the error bars in this bar plot. Thank you very much.
data
yield <- structure(
list(
N_rate = structure(
c(
4L,
5L,
3L,
1L,
2L,
5L,
3L,
4L,
1L,
2L,
1L,
4L,
5L,
2L,
3L,
1L,
2L,
4L,
3L,
5L,
1L,
3L,
2L,
4L,
5L,
3L,
5L,
4L,
2L,
1L,
5L,
1L,
2L,
4L,
3L,
4L,
3L,
1L,
2L,
5L,
2L,
5L,
3L,
1L,
4L,
3L,
5L,
1L,
2L,
4L,
3L,
5L,
1L,
4L,
2L,
2L,
5L,
1L,
4L,
3L
),
levels = c("0", "45", "90", "179", "269"),
class = "factor"
),
History = structure(
c(
2L,
2L,
2L,
2L,
2L,
3L,
3L,
3L,
3L,
3L,
1L,
1L,
1L,
1L,
1L,
1L,
1L,
1L,
1L,
1L,
2L,
2L,
2L,
2L,
2L,
3L,
3L,
3L,
3L,
3L,
3L,
3L,
3L,
3L,
3L,
1L,
1L,
1L,
1L,
1L,
2L,
2L,
2L,
2L,
2L,
2L,
2L,
2L,
2L,
2L,
3L,
3L,
3L,
3L,
3L,
1L,
1L,
1L,
1L,
1L
),
levels = c("fallow", "corn", "soybean"),
class = "factor"
),
Rep = structure(
c(
1L,
1L,
1L,
1L,
1L,
1L,
1L,
1L,
1L,
1L,
1L,
1L,
1L,
1L,
1L,
2L,
2L,
2L,
2L,
2L,
2L,
2L,
2L,
2L,
2L,
2L,
2L,
2L,
2L,
2L,
3L,
3L,
3L,
3L,
3L,
3L,
3L,
3L,
3L,
3L,
3L,
3L,
3L,
3L,
3L,
4L,
4L,
4L,
4L,
4L,
4L,
4L,
4L,
4L,
4L,
4L,
4L,
4L,
4L,
4L
),
levels = c("1", "2", "3", "4"),
class = "factor"
),
`Yield at std moisture (kg/ha)` = c(
12346.9249590533,
11860.0460239527,
6286.35275360947,
2793.90516411834,
5127.27805576331,
12312.7660459172,
12086.4480249941,
12772.9435023905,
7781.93368615385,
10711.1602139645,
7813.41363351479,
12249.8439422485,
13803.4656340828,
7778.30574504142,
11177.9322092308,
4926.47739379882,
7942.4952744142,
11233.1764305799,
10579.7313287574,
12547.0432825562,
3660.44141822485,
10565.0201115266,
7381.04619323077,
10584.2536581302,
12692.5976236686,
11582.7898622485,
12617.2233679527,
13225.3150068639,
11339.0543647811,
9553.99606381065,
12957.7648473373,
8697.03144369231,
9722.50007176331,
12775.756836355,
12011.991252071,
13313.2421907692,
11158.0877072663,
7208.26549775148,
8189.39052383432,
14138.8465351953,
8795.41415233136,
13314.8147184852,
10684.0892228166,
6489.22772449704,
12286.5138609231,
10587.1551712189,
12207.7824999763,
4808.05912842604,
8768.34316118343,
11741.9321865089,
11842.0952736568,
12872.0694350769,
9509.94219323077,
13085.1710848757,
11423.857488284,
8773.69689372781,
12733.0655431953,
7236.61298669823,
12536.2749318817,
11056.5389481657
)
),
row.names = c(NA, -60L),
class = c("tbl_df", "tbl", "data.frame")
)
Convert character to factor
yield$History <- factor(yield$History,levels = c('fallow','corn','soybean'))
yield$N_rate <- factor(yield$N_rate)
yield$Rep <- factor(yield$Rep)
Fit a linear model
mod_plot <- lm(Yield at std moisture (kg/ha)~Rep+History*N_rate,data=yield)
Letters <- agricolae::LSD.test(y = mod_plot,trt = c('N_rate','History'),console = T,group = T)
LSD_table <- Letters$groups # Extracts means and groups
Add proper column names for interaction terms
LSD_table <- LSD_table %>%
tibble::rownames_to_column(var = "Interaction_Term") %>% # Convert row names to a column
tidyr::separate(Interaction_Term, into = c("N_rate", "History"), sep = ":", fill = "right")
Print formatted table
print(LSD_table)
First, calculate summary stats (mean and SE)
summary_data <- yield %>%
group_by(N_rate, History) %>%
summarise(
mean = mean(`Yield at std moisture (kg/ha)`, na.rm = TRUE),
se = sd(`Yield at std moisture (kg/ha)`, na.rm = TRUE) / sqrt(n()),
.groups = "drop"
)
Merge with LSD table for labels
label_data <- summary_data %>%
left_join(LSD_table, by = c("N_rate", "History")) %>%
mutate(label_y = mean + se) # place label at center of error bar top
Final plot
p1 <- ggplot(data = yield, aes(x = N_rate, y = `Yield at std moisture (kg/ha)`, fill = History)) +
# Error bars
stat_summary(
geom = 'errorbar',
position = position_dodge(.9),
width = .3,
fun.data = 'mean_se', col = 'black'
) +
# Bars
stat_summary(
geom = 'col',
position = position_dodge(.9),
fun = mean
) +
scale_fill_brewer(palette = 'Dark2') +
scale_y_continuous(expand = expansion(mult = c(0, 0.2), add = c(0, 0.4))) +
ylab(expression(bold(Yield ~ (kg ~ ha ^ -1)))) +
xlab(expression(bold(Nitrogen ~rate ~ (kg ~ ha ^ -1)))) +
scale_fill_manual(name = '', values = c('#7570B3','#D95F02','#1B9E77')) +
# Text labels exactly at the top of error bars
geom_text(
data = label_data,
aes(
x = N_rate,
y = label_y,
label = groups,
fill = History
),
position = position_dodge(.9),
show.legend = FALSE,
vjust = -0.9,
fontface='bold'
) +
theme_test() +
theme(
legend.position = c(0.1, 0.9),
axis.title = element_text(
family = 'serif',
face = 'bold',
colour = 'black',
size = 16
),
axis.text = element_text(
family = 'serif',
face = 'bold',
colour = 'black',
size = 16
),
legend.text = element_text(
family = 'serif',
face = 'bold',
colour = 'black',
size = 16
)
)
p1