Using the shortest.paths function we get the shortest path from a graph. Now, I want to limit the length of the shortest path.
For example, when I am running the below code, I am getting all the shortest paths from a vertex to any vertex.
df <- read.csv("~/data.csv")
g1 <- df
graph1 <- graph_from_data_frame(g1, directed = FALSE)
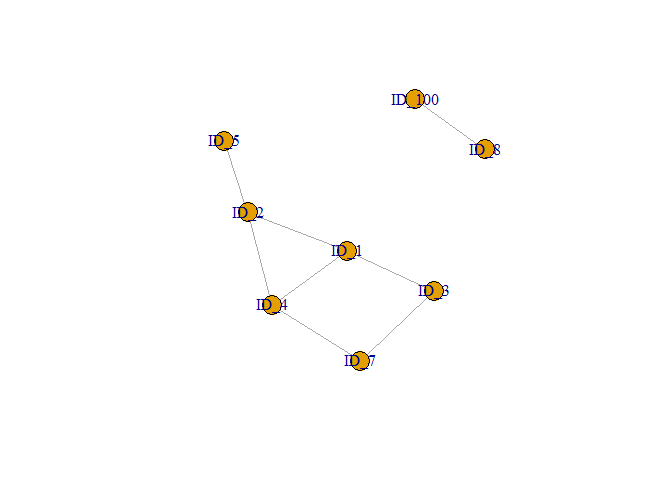
plot(graph1, vertex.label = V(graph1)$name)
mat <- shortest.paths(graph1)
The output I am getting
ID_1 ID_2 ID_3 ID_4 ID_8 ID_5 ID_7 ID_100
ID_1 0 1 1 1 Inf 2 2 Inf
ID_2 1 0 2 1 Inf 1 2 Inf
ID_3 1 2 0 2 Inf 3 1 Inf
ID_4 1 1 2 0 Inf 2 1 Inf
ID_8 Inf Inf Inf Inf 0 Inf Inf 1
ID_5 2 1 3 2 Inf 0 3 Inf
ID_7 2 2 1 1 Inf 3 0 Inf
ID_100 Inf Inf Inf Inf 1 Inf Inf 0
But, I want to keep only (say) the path length is 3 and the other will be 0 or Inf. Actually, I do not need other except (path length =3).
Moreover, I want the sum of the path weight not only the number of the path. I thought I Can do this just by changing only one line
mat <- shortest.paths(graph1, weights=E(graph1)$weight)
But, how can limit the path length?
Reproducible Data
structure(list(nodeA = structure(c(1L, 1L, 1L, 2L, 2L, 3L, 4L,
5L), .Label = c("ID_1", "ID_2", "ID_3", "ID_4", "ID_8"), class = "factor"),
nodeB = structure(c(2L, 3L, 4L, 5L, 4L, 6L, 6L, 1L), .Label = c("ID_100",
"ID_2", "ID_3", "ID_4", "ID_5", "ID_7"), class = "factor"),
weight = c(0.5, 0.77, 0.5, 0.9, 0.44, 0.32, 0.45, 0.543)), class = "data.frame", row.names = c(NA,
-8L))