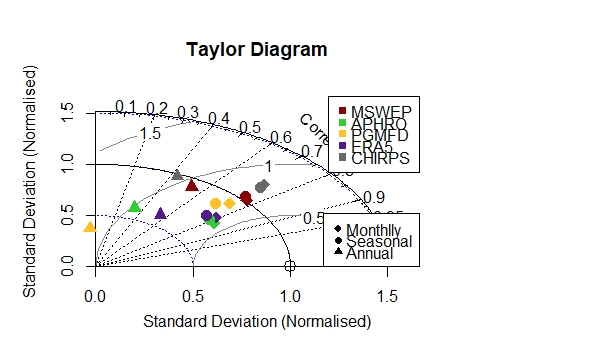
I have developed taylor diagram and trying to change color and type of lines but unable to do it. Secondly, legends are overlaying on diagram Kindly help me to resolve these issues. The diagram and code are given below:
library(plotrix)
data = read.csv("taylor-GOF-upstream.csv")
taylor.diagram(as.vector(data$OBS),as.vector(data$MSWEP),add=FALSE,col="red4",
pch=18,pos.cor=TRUE,xlab="Standard Deviation (Normalised)",
ylab="Standard Deviation (Normalised)",
show.gamma=TRUE,ngamma=5,gamma.col="honeydew4",sd.arcs=1,ref.sd=TRUE,
grad.corr.lines=c(0.2,0.4,0.6,0.8, 0.9, 0.95),
pcex=1.7,cex.axis=1,normalize=TRUE,mar=c(5,5,5,10),
lwd=10,font=3,lty=3)
taylor.diagram(as.vector(data$OBS),as.vector(data$APHRO),add=TRUE,col="limegreen",
pch=18,pos.cor=TRUE,xlab="Standard Deviation (Normalised)",
ylab="Standard Deviation (Normalised)",
show.gamma=TRUE,ngamma=5,gamma.col="honeydew4",sd.arcs=1,ref.sd=TRUE,
grad.corr.lines=c(0.2,0.4,0.6,0.8, 0.9, 0.95),
pcex=1.7,cex.axis=1,normalize=TRUE,mar=c(5,6,5,10),
lwd=10,font=3,lty=3)
taylor.diagram(as.vector(data$OBS),as.vector(data$PGMFD),add=TRUE,col="goldenrod1",
pch=18,pos.cor=TRUE,xlab="Standard Deviation (Normalised)",
ylab="Standard Deviation (Normalised)",
show.gamma=TRUE,ngamma=5,gamma.col="honeydew4",sd.arcs=1,ref.sd=TRUE,
grad.corr.lines=c(0.2,0.4,0.6,0.8, 0.9, 0.95),
pcex=1.7,cex.axis=1,normalize=TRUE,mar=c(5,6,5,10),
lwd=10,font=3,lty=3)
taylor.diagram(as.vector(data$OBS),as.vector(data$ERA5),add=TRUE,col="purple4",
pch=18,pos.cor=TRUE,xlab="Standard Deviation (Normalised)",
ylab="Standard Deviation (Normalised)",
show.gamma=TRUE,ngamma=5,gamma.col="honeydew4",sd.arcs=1,ref.sd=TRUE,
grad.corr.lines=c(0.2,0.4,0.6,0.8, 0.9, 0.95),
pcex=1.7,cex.axis=1,normalize=TRUE,mar=c(5,6,5,10),
lwd=10,font=3,lty=3)
taylor.diagram(as.vector(data$OBS),as.vector(data$CHIRPS),add=TRUE,col="dimgrey",
pch=18,pos.cor=TRUE,xlab="Standard Deviation (Normalised)",
ylab="Standard Deviation (Normalised)",
show.gamma=TRUE,ngamma=5,gamma.col="honeydew4",sd.arcs=1,ref.sd=TRUE,
grad.corr.lines=c(0.2,0.4,0.6,0.8, 0.9, 0.95),
pcex=1.7,cex.axis=1,normalize=TRUE,mar=c(5,6,5,10),
lwd=10,font=3,lty=3)
taylor.diagram(as.vector(data$OBS.S),as.vector(data$MSWEP.S),add=TRUE,col="red4",
pch=16,pos.cor=TRUE,xlab="Standard Deviation (Normalised)",
ylab="Standard Deviation (Normalised)",
show.gamma=TRUE,ngamma=5,gamma.col="honeydew4",sd.arcs=1,ref.sd=TRUE,
grad.corr.lines=c(0.2,0.4,0.6,0.8, 0.9, 0.95),
pcex=1.5,cex.axis=1,normalize=TRUE,mar=c(5,6,5,10),
lwd=10,font=3,lty=3)
taylor.diagram(as.vector(data$OBS.S),as.vector(data$APHRO.S),add=TRUE,col="limegreen",
pch=16,pos.cor=TRUE,xlab="Standard Deviation (Normalised)",
ylab="Standard Deviation (Normalised)",
show.gamma=TRUE,ngamma=5,gamma.col="honeydew4",sd.arcs=1,ref.sd=TRUE,
grad.corr.lines=c(0.2,0.4,0.6,0.8, 0.9, 0.95),
pcex=1.5,cex.axis=1,normalize=TRUE,mar=c(5,6,5,10),
lwd=10,font=3,lty=3)
taylor.diagram(as.vector(data$OBS.S),as.vector(data$PGMFD.S),add=TRUE,col="goldenrod1",
pch=16,pos.cor=TRUE,xlab="Standard Deviation (Normalised)",
ylab="Standard Deviation (Normalised)",
show.gamma=TRUE,ngamma=5,gamma.col="honeydew4",sd.arcs=1,ref.sd=TRUE,
grad.corr.lines=c(0.2,0.4,0.6,0.8, 0.9, 0.95),
pcex=1.5,cex.axis=1,normalize=TRUE,mar=c(5,6,5,10),
lwd=10,font=3,lty=3)
taylor.diagram(as.vector(data$OBS.S),as.vector(data$ERA5.S),add=TRUE,col="purple4",
pch=16,pos.cor=TRUE,xlab="Standard Deviation (Normalised)",
ylab="Standard Deviation (Normalised)",
show.gamma=TRUE,ngamma=5,gamma.col="honeydew4",sd.arcs=1,ref.sd=TRUE,
grad.corr.lines=c(0.2,0.4,0.6,0.8, 0.9, 0.95),
pcex=1.5,cex.axis=1,normalize=TRUE,mar=c(5,6,5,10),
lwd=10,font=3,lty=3)
taylor.diagram(as.vector(data$OBS.S),as.vector(data$CHIRPS.S),add=TRUE,col="dimgrey",
pch=16,pos.cor=TRUE,xlab="Standard Deviation (Normalised)",
ylab="Standard Deviation (Normalised)",
show.gamma=TRUE,ngamma=5,gamma.col="honeydew4",sd.arcs=1,ref.sd=TRUE,
grad.corr.lines=c(0.2,0.4,0.6,0.8, 0.9, 0.95),
pcex=1.5,cex.axis=1,normalize=TRUE,mar=c(5,6,5,10),
lwd=10,font=3,lty=3)
taylor.diagram(as.vector(data$OBS.A),as.vector(data$MSWEP.A),add=TRUE,col="red4",
pch=17,pos.cor=TRUE,xlab="Standard Deviation (Normalised)",
ylab="Standard Deviation (Normalised)",
show.gamma=TRUE,ngamma=5,gamma.col="honeydew4",sd.arcs=1,ref.sd=TRUE,
grad.corr.lines=c(0.2,0.4,0.6,0.8, 0.9, 0.95),
pcex=1.5,cex.axis=1,normalize=TRUE,mar=c(5,6,5,10),
lwd=10,font=3,lty=3)
taylor.diagram(as.vector(data$OBS.A),as.vector(data$APHRO.A),add=TRUE,col="limegreen",
pch=17,pos.cor=TRUE,xlab="Standard Deviation (Normalised)",
ylab="Standard Deviation (Normalised)",
show.gamma=TRUE,ngamma=5,gamma.col="honeydew4",sd.arcs=1,ref.sd=TRUE,
grad.corr.lines=c(0.2,0.4,0.6,0.8, 0.9, 0.95),
pcex=1.5,cex.axis=1,normalize=TRUE,mar=c(5,6,5,10),
lwd=10,font=3,lty=3)
taylor.diagram(as.vector(data$OBS.A),as.vector(data$PGMFD.A),add=TRUE,col="goldenrod1",
pch=17,pos.cor=TRUE,xlab="Standard Deviation (Normalised)",
ylab="Standard Deviation (Normalised)",
show.gamma=TRUE,ngamma=5,gamma.col="honeydew4",sd.arcs=1,ref.sd=TRUE,
grad.corr.lines=c(0.2,0.4,0.6,0.8, 0.9, 0.95),
pcex=1.5,cex.axis=1,normalize=TRUE,mar=c(5,6,5,10),
lwd=10,font=3,lty=3)
taylor.diagram(as.vector(data$OBS.A),as.vector(data$ERA5.A),add=TRUE,col="purple4",
pch=17,pos.cor=TRUE,xlab="Standard Deviation (Normalised)",
ylab="Standard Deviation (Normalised)",
show.gamma=TRUE,ngamma=5,gamma.col="honeydew4",sd.arcs=1,ref.sd=TRUE,
grad.corr.lines=c(0.2,0.4,0.6,0.8, 0.9,0.95),
pcex=1.5,cex.axis=1,normalize=TRUE,mar=c(5,6,5,10),
lwd=10,font=3,lty=3)
taylor.diagram(as.vector(data$OBS.A),as.vector(data$CHIRPS.A),add=TRUE,col="dimgrey",
pch=17,pos.cor=TRUE,xlab="Standard Deviation (Normalised)",
ylab="Standard Deviation (Normalised)",
show.gamma=TRUE,ngamma=5,gamma.col="honeydew4",sd.arcs=1,ref.sd=TRUE,
grad.corr.lines=c(0.2,0.4,0.6,0.8, 0.9, 0.95),
pcex=1.5,cex.axis=1,normalize=TRUE,mar=c(5,6,5,10),
lwd=10,font=3,lty=3)
legend("bottomright", # << THIS IS THE HACKISH PART
legend=c("Monthlly","Seasonal","Annual"),
col=c(rep("black",3)), pch=c(18,16,17),
xjust = 1, yjust = 1, x.intersp = 0.6, y.intersp = 0.6, text.font =0.8, ncol=1, title.adj = 0.4)
legend("topright", # << THIS IS THE HACKISH PART
legend=c("MSWEP", "APHRO", "PGMFD","ERA5","CHIRPS"),
col=c('red4','limegreen','goldenrod1','purple4','dimgrey'), pch=c(15,15,15,15,15),
xjust = 1, yjust = 1, x.intersp = 0.6, y.intersp = 0.6, text.font =0.8, ncol=1, title.adj = 0.4)