I would like to manually generate a ggplot for Johnson-Neyman technique output given by SPSS PROCESS macro, in this style:
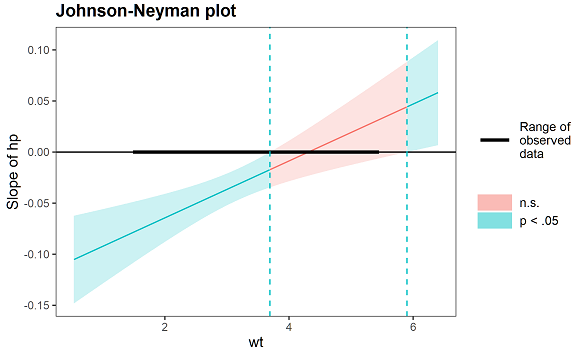
However, I have been unable to figure out how to:
- Colour the space between confidence interval bands and the line for regions of significance and non-significance differently
- Create a legend inside the graph for the three lines: upper 95% CI, Effect, lower 95% CI. I feel text annotations are quite unsightly.
- Label the point at which my moderator becomes significant with 2 d.p. but to have all other intervals without any decimal places.
Here is what I have at the moment, with the green dotted lines pointing to the point where my moderator crossed into significance. In my case, the bands to the left of x = 3.2381 will be n.s. (red) and everything to the right is significant (green).
# The SPSS PROCESS J-N output
jnplot <- tibble::tribble(
~chb_MEAN, ~Effect, ~se, ~t, ~p, ~LLCI, ~ULCI,
1, -0.5887, 1.0072, -0.5845, 0.5593, -2.5708, 1.3934,
1.2583, -0.4308, 0.923, -0.4667, 0.6411, -2.2472, 1.3856,
1.5167, -0.2728, 0.8402, -0.3247, 0.7456, -1.9263, 1.3807,
1.775, -0.1149, 0.7593, -0.1513, 0.8799, -1.6092, 1.3794,
2.0333, 0.0431, 0.681, 0.0632, 0.9496, -1.2972, 1.3833,
2.2917, 0.201, 0.6063, 0.3315, 0.7405, -0.9922, 1.3942,
2.55, 0.3589, 0.5367, 0.6688, 0.5041, -0.6972, 1.415,
2.8083, 0.5169, 0.4743, 1.0897, 0.2767, -0.4166, 1.4503,
3.0667, 0.6748, 0.4226, 1.5968, 0.1114, -0.1568, 1.5064,
3.2381, 0.7796, 0.3962, 1.9679, 0.05, 0, 1.5592,
3.325, 0.8328, 0.3857, 2.1591, 0.0316, 0.0737, 1.5918,
3.5833, 0.9907, 0.3681, 2.6911, 0.0075, 0.2662, 1.7152,
3.8417, 1.1486, 0.3727, 3.0821, 0.0022, 0.4152, 1.882,
4.1, 1.3066, 0.3986, 3.2783, 0.0012, 0.5223, 2.0909,
4.3583, 1.4645, 0.442, 3.3132, 0.001, 0.5946, 2.3344,
4.6167, 1.6224, 0.4985, 3.2545, 0.0013, 0.6414, 2.6035,
4.875, 1.7804, 0.5641, 3.1559, 0.0018, 0.6702, 2.8906,
5.1333, 1.9383, 0.6361, 3.0474, 0.0025, 0.6866, 3.19,
5.3917, 2.0963, 0.7124, 2.9426, 0.0035, 0.6944, 3.4982,
5.65, 2.2542, 0.7918, 2.8468, 0.0047, 0.6959, 3.8125,
5.9083, 2.4121, 0.8736, 2.7613, 0.0061, 0.6931, 4.1312,
6.1667, 2.5701, 0.957, 2.6856, 0.0076, 0.6868, 4.4533
)
# compute CIs
errorbands <- jnplot %>% select(chb_MEAN, se, Effect) %>%
mutate(lowerci = chb_MEAN-se,
upperci = chb_MEAN+se)
ggplot(errorbands, aes(x = chb_MEAN, y = Effect)) +
geom_line(aes(x = chb_MEAN), size = 1, color = "#F8766D") +
geom_line(aes(x = upperci), lty = 2, size = 0.6, color = "grey") +
geom_line(aes(x = lowerci), lty = 2, size = 0.6, color = "grey") +
#geom_ribbon(aes(ymin=lowerci, ymax=upperci, x=chb_MEAN, fill = "red"), alpha = 0.3)+
# vertical line
geom_segment(aes(x = 3.2381, y = 0.7796,
xend = 3.2381, yend = -Inf),
linetype = "dashed", col = "green") +
# horizontal line
geom_segment(aes(x = 3.2381, y = 0.7796,
xend = -Inf, yend = 0.7796),
linetype = "dashed", col = "green") +
annotate("text", label = "Upper 95% CI", x = 4.4, y = 2.5, size = 3, colour = "black") +
annotate("text", label = "Lower 95% CI", x = 6.9, y = 2.1, size = 3, colour = "black") +
scale_x_continuous(breaks = c(0, 2, 3.2381, 4, 6)) + # does not work
labs(x = "CHB Mean", y = "Effect") +
theme_bw() +
theme(panel.grid.major = element_blank(),
panel.grid.minor = element_blank())
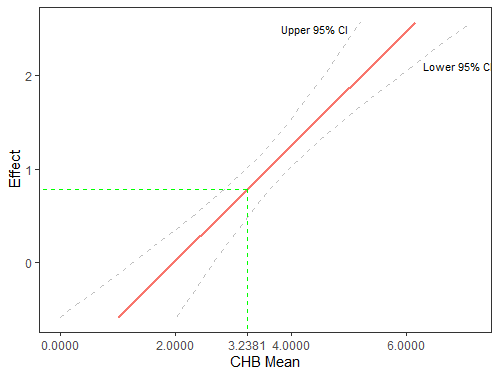
Thanks for any help!