Hello,
I am trying to work on a for loop to make running a function I've developed more efficient.
However, when I put it in a for loop, it is overwriting columns that it should not be and returning incorrect results. Can someone please tell me why is it returning these incorrect results and what I can do to fix it? I will upload the relevant code below.
Edit: My code and dataframes are now in preformatted text after using datapasta.
#2.) Outlier_Hunter Function
#Function to Generate the Outlier table
#Outlier Hunter function takes 4 arguments: the dataset, column/variable of interest,
#Q1 and Q3. Q1 and Q3 are stored in the results of Quartile_Hunter.
#Input ex: MiSeq_Bord_final_report0, Avg_Trim_Cov, MiSeq_Bord_Quartiles_ATC$First_Quartile[1], MiSeq_Bord_Quartiles_ATC$Third_Quartile[1]
#Usage ex: Outlier_Hunter(MiSeq_Bord_final_report0, Avg_Trim_Cov,
#MiSeq_Bord_Quartiles_ATC$First_Quartile[1], MiSeq_Bord_Quartiles_ATC$Third_Quartile[1])
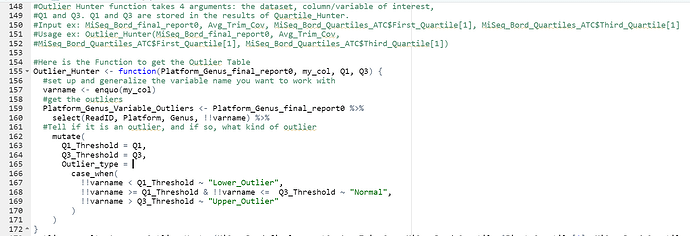
#Here is the Function to get the Outlier Table
Outlier_Hunter <- function(Platform_Genus_final_report0, my_col, Q1, Q3) {
#set up and generalize the variable name you want to work with
varname <- enquo(my_col)
print(varname)
#get the outliers
Platform_Genus_Variable_Outliers <- Platform_Genus_final_report0 %>%
select(ReadID, Platform, Genus, !!varname) %>%
#Tell if it is an outlier, and if so, what kind of outlier
mutate(
Q1_Threshold = Q1,
Q3_Threshold = Q3,
Outlier_type =
case_when(
!!varname < Q1_Threshold ~ "Lower_Outlier",
!!varname >= Q1_Threshold & !!varname <= Q3_Threshold ~ "Normal",
!!varname > Q3_Threshold ~ "Upper_Outlier"
)
)
}
#Execution for 1 variable
outlier_results_1var <- Outlier_Hunter(MiSeq_Bord_final_report0, Avg_Trim_Cov,
MiSeq_Bord_Quartiles$First_Quartile[1], MiSeq_Bord_Quartiles$Third_Quartile[1])
#Now do it with a for loop
col_var_outliers <- row.names(MiSeq_Bord_Quartiles)
#col_var_outliers <- c("Avg_Trim_Cov", "S2_Total_Read_Pairs_Processed")
#change line above to change input of variables few into Outlier Hunter Function
outlier_list_MiSeq_Bord <- list()
i <- 0
for (y in col_var_outliers) {
i <- i + 1
#outlier_results0 <- Outlier_Hunter(MiSeq_Bord_final_report0, y, MiSeq_Bord_Quartiles[y, "First_Quartile"],
#MiSeq_Bord_Quartiles[y, "Third_Quartile"])
outlier_results0 <- Outlier_Hunter(MiSeq_Bord_final_report0, y, MiSeq_Bord_Quartiles$First_Quartile[i],
MiSeq_Bord_Quartiles$Third_Quartile[i])
#print(head(MiSeq_Bord_final_report0), y, MiSeq_Bord_Quartiles$First_Quartile[i],
# MiSeq_Bord_Quartiles$Third_Quartile[i])
outlier_results1 <- outlier_results0
colnames(outlier_results1)[5:7] <- paste0(y, "_", colnames(outlier_results1[, c(5:7)]), sep = "")
outlier_list_MiSeq_Bord[[i]] <- outlier_results1
}
MiSeq_Bord_Outliers_table0 <- reduce(outlier_list_MiSeq_Bord, left_join, by = c("ReadID", "Platform", "Genus"))
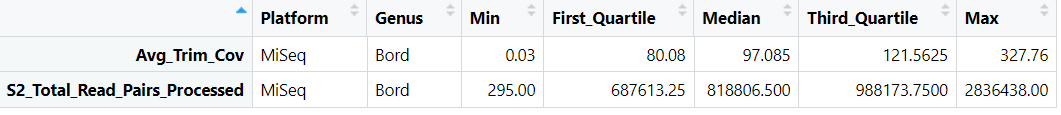
#MiSeq_Bord_Quartiles entries
datapasta::df_paste(head(MiSeq_Bord_Quartiles, 5))
data.frame(
stringsAsFactors = FALSE,
row.names = c("Avg_Trim_Cov", "S2_Total_Read_Pairs_Processed"),
Platform = c("MiSeq", "MiSeq"),
Genus = c("Bord", "Bord"),
Min = c(0.03, 295),
First_Quartile = c(80.08, 687613.25),
Median = c(97.085, 818806.5),
Third_Quartile = c(121.5625, 988173.75),
Max = c(327.76, 2836438)
)
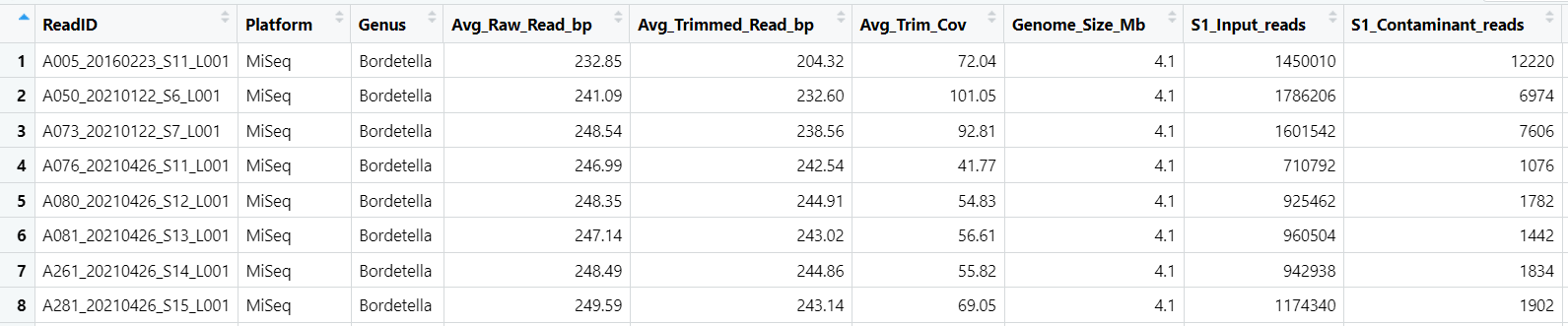
#dataset entry
datapasta::df_paste(head(MiSeq_Bord_final_report0, 5))
data.frame(
stringsAsFactors = FALSE,
ReadID = c("A005_20160223_S11_L001","A050_20210122_S6_L001",
"A073_20210122_S7_L001",
"A076_20210426_S11_L001",
"A080_20210426_S12_L001"),
Platform = c("MiSeq","MiSeq",
"MiSeq","MiSeq","MiSeq"),
Genus = c("Bordetella",
"Bordetella","Bordetella",
"Bordetella","Bordetella"),
Avg_Raw_Read_bp = c(232.85,241.09,
248.54,246.99,248.35),
Avg_Trimmed_Read_bp = c(204.32,232.6,
238.56,242.54,244.91),
Avg_Trim_Cov = c(72.04,101.05,
92.81,41.77,54.83),
Genome_Size_Mb = c(4.1, 4.1, 4.1, 4.1, 4.1),
S1_Input_reads = c(1450010L,
1786206L,1601542L,710792L,925462L),
S1_Contaminant_reads = c(12220L,6974L,
7606L,1076L,1782L),
S1_Total_reads_removed = c(12220L,6974L,
7606L,1076L,1782L),
S1_Result_reads = c(1437790L,
1779232L,1593936L,709716L,923680L),
S2_Read_Pairs_Written = c(712776L,882301L,
790675L,352508L,459215L),
S2_Total_Read_Pairs_Processed = c(718895L,889616L,
796968L,354858L,461840L)
)data.frame(
stringsAsFactors = FALSE,
ReadID = c("A005_20160223_S11_L001","A050_20210122_S6_L001",
"A073_20210122_S7_L001",
"A076_20210426_S11_L001","A080_20210426_S12_L001"),
Platform = c("MiSeq", "MiSeq", "MiSeq", "MiSeq", "MiSeq"),
Genus = c("Bordetella","Bordetella","Bordetella",
"Bordetella","Bordetella"),
Avg_Raw_Read_bp = c(232.85, 241.09, 248.54, 246.99, 248.35),
Avg_Trimmed_Read_bp = c(204.32, 232.6, 238.56, 242.54, 244.91),
Avg_Trim_Cov = c(72.04, 101.05, 92.81, 41.77, 54.83),
Genome_Size_Mb = c(4.1, 4.1, 4.1, 4.1, 4.1),
S1_Input_reads = c(1450010L,1786206L,1601542L,710792L,925462L),
S1_Contaminant_reads = c(12220L, 6974L, 7606L, 1076L, 1782L),
S1_Total_reads_removed = c(12220L, 6974L, 7606L, 1076L, 1782L),
S1_Result_reads = c(1437790L,1779232L,1593936L,709716L,923680L),
S2_Read_Pairs_Written = c(712776L, 882301L, 790675L, 352508L, 459215L),
S2_Total_Read_Pairs_Processed = c(718895L, 889616L, 796968L, 354858L, 461840L)
)
#the columns containing label Outlier_type is where the code goes wrong
datapasta::df_paste(head(MiSeq_Bord_Outliers_table0, 5))
data.frame(
stringsAsFactors = FALSE,
ReadID = c("A005_20160223_S11_L001",
"A050_20210122_S6_L001",
"A073_20210122_S7_L001","A076_20210426_S11_L001",
"A080_20210426_S12_L001"),
Platform = c("MiSeq",
"MiSeq","MiSeq","MiSeq",
"MiSeq"),
Genus = c("Bordetella","Bordetella","Bordetella",
"Bordetella","Bordetella"),
Avg_Trim_Cov = c(72.04,
101.05,92.81,41.77,54.83),
Avg_Trim_Cov_Q1_Threshold = c(80.08,
80.08,80.08,80.08,80.08),
Avg_Trim_Cov_Q3_Threshold = c(121.5625,
121.5625,121.5625,121.5625,
121.5625),
Avg_Trim_Cov_Outlier_type = c("Upper_Outlier","Upper_Outlier",
"Upper_Outlier","Upper_Outlier",
"Upper_Outlier"),
S2_Total_Read_Pairs_Processed = c(718895L,
889616L,796968L,354858L,
461840L),
S2_Total_Read_Pairs_Processed_Q1_Threshold = c(687613.25,
687613.25,687613.25,
687613.25,687613.25),
S2_Total_Read_Pairs_Processed_Q3_Threshold = c(988173.75,
988173.75,988173.75,
988173.75,988173.75),
S2_Total_Read_Pairs_Processed_Outlier_type = c("Upper_Outlier","Upper_Outlier",
"Upper_Outlier","Upper_Outlier",
"Upper_Outlier")
)