Hi @paulogil2010! Welcome!
Thanks for providing the data, which made it easy for me to see what your problem was 
So there are two things going awry in the code you posted — but the good news is that the findpeaks() part is working just fine! 
The first issue is with how the CSV file gets imported. Note that I'm assuming here that you are importing the exact same CSV file you posted online. If that's not the case, this part might not apply to you.
library(pracma)
# Btw, here's an example of one way to write a CSV file import when constructing a
# self-contained reproducible example that somebody else can easily run
data_url <- "https://raw.githubusercontent.com/paulogil2010/hello-world/master/wave_vector.csv"
data_path <- file.path(tempdir(), "wave_vector.csv")
download.file(data_url, data_path)
data <- read.csv(data_path)
str(data)
#> 'data.frame': 1000 obs. of 2 variables:
#> $ X: int 1 2 3 4 5 6 7 8 9 10 ...
#> $ x: num 290 309 329 351 372 ...
There are two columns here, and one of them has been (confusingly!) given the automatic name of capital X. Where did this X column come from? Here are the first three lines of the CSV file:
cat(readLines(data_path, n = 3), sep = "\n")
#> "","x"
#> "1",289.8887
#> "2",308.60833
The file has row names, but read.csv() is interpreting them as a column of data that is missing a name. (To force them to be read in as row names in this case, you'd use read.csv("wave_vector.csv", row.names = 1)).
Flattening a data frame with two columns results in the column of row names being appended to the beginning of the column of data, which is not what you want!
str(purrr::flatten_dbl(data))
#> num [1:2000] 1 2 3 4 5 6 7 8 9 10 ...
But flattening is overkill here — you can just pull the vector out of the data frame using dollar-sign or double-bracket indexing (or dplyr offers the pull() function to do the same thing).
wave_vector <- data$x
str(wave_vector)
#> num [1:1000] 290 309 329 351 372 ...
# Finally! Find some peaks! :-)
peaks <- findpeaks(wave_vector, minpeakdistance = 100)
Now we encounter problem #2: peaks is a matrix, with column definitions given in the documentation:
The first column gives the height, the second the position/index where the maximum is reached, the third and fourth the indices of where the peak begins and ends — in the sense of where the pattern starts and ends.
peaks
#> [,1] [,2] [,3] [,4]
#> [1,] 486.8127 977 955 997
#> [2,] 481.9355 15 1 41
#> [3,] 450.8867 135 116 159
#> [4,] 448.6920 858 838 882
#> [5,] 448.6058 375 356 399
#> [6,] 448.1287 495 475 519
#> [7,] 447.9364 617 598 642
#> [8,] 446.8559 255 235 279
#> [9,] 444.3782 739 719 763
So to get the plot I think you’re looking for, you need to grab the values from the second column of peaks
plot(wave_vector, type = 'l')
points(peaks[, 2], wave_vector[peaks[, 2]])
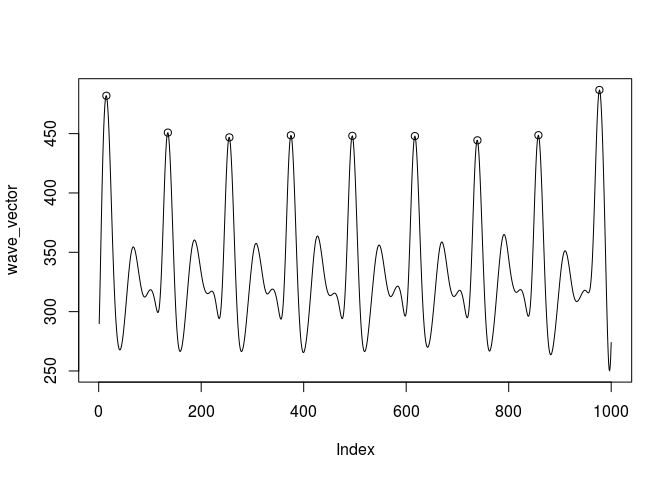
Created on 2018-11-01 by the reprex package (v0.2.1)


 ). By including the output along with the code and enforcing a decent amount of self-contained-ness,
). By including the output along with the code and enforcing a decent amount of self-contained-ness,