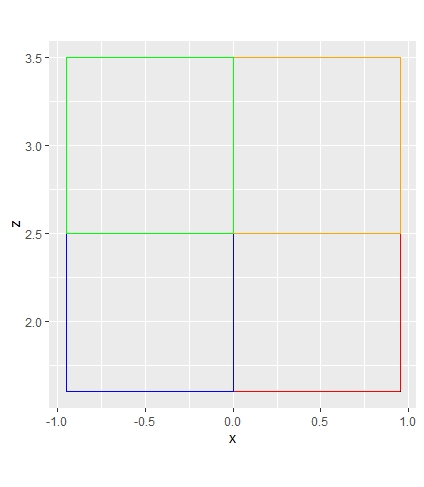
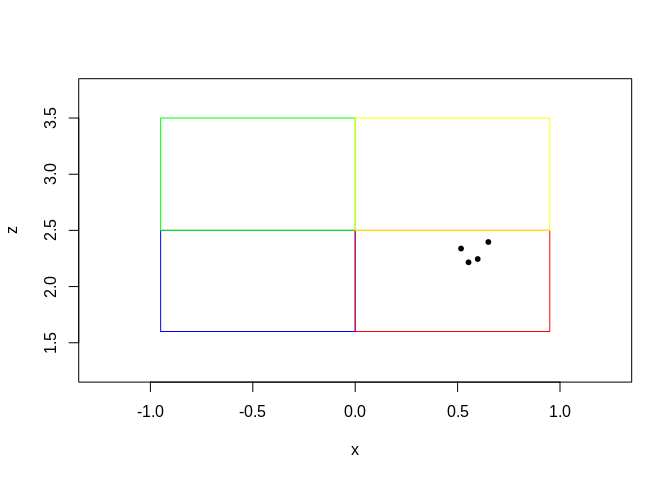
Hi, I was wondering if there is a way to filter my data by a specific region of points. So, in the picture I have below, I would like to see just the points that are in the red region (down_in). Thank you!
yelich <- data.frame(
plate_x = c(-0.4895, -0.5197, 0.5169, 1.1077, 1.3299, 0.1347, 1.3979,
-0.2142, 0.5982, -0.5614, 0.6501, 0.3105, 1.2385, 0.2096,
0.48, 0.5458, -0.7775, -0.9248, 1.4628, 1.2406, -0.1379, -0.2188,
-0.7861, -2.4055, -0.2201, -0.9993, 0.5534, 1.8461, -0.2148,
-0.0624),
plate_z = c(3.8125, 2.6877, 2.3384, 1.8482, 1.2539, 1.4041, 2.7248,
2.2688, 2.2451, 1.7012, 2.3968, 0.9775, 2.7827, 3.2931,
0.6966, 3.6894, 2.3352, 1.8292, 2.5673, 2.8705, 1.4959, 1.9526,
-0.6215, 1.5998, 1.9236, 0.7451, 2.2158, 2.189, -0.0983, 1.4302)
)
down_in_x <- c(.95,0, 0, .95, .95)
down_in_y <- c(1.6, 1.6, 2.5, 2.5, 1.6)
down_in <- data.frame(down_in_x, down_in_y)
down_out_x <- c(-.95,-.95, 0, 0, -.95)
down_out_y <- c(1.6, 2.5, 2.5, 1.6, 1.6)
down_out <- data.frame(down_out_x, down_out_y)
up_in_x <- c(0,0,.95,.95,0)
up_in_y <- c(2.5,3.5,3.5,2.5,2.5)
up_in <- data.frame(up_in_x, up_in_y)
up_out_x <- c(0,0,-.95, -.95, 0)
up_out_y <- c(2.5,3.5,3.5, 2.5, 2.5)
up_out <- data.frame(up_out_x, up_out_y)