rm(list = ls())
setwd(dirname(rstudioapi::getActiveDocumentContext()$path))
#> Error: RStudio not running
getwd()
#> [1] "C:/Users/Angela/AppData/Local/Temp/RtmpUZ4zh3/reprex-26305ae4190-awful-dog"
#load required packages
library(mc2d)
#> Loading required package: mvtnorm
#>
#> Attaching package: 'mc2d'
#> The following objects are masked from 'package:base':
#>
#> pmax, pmin
library(gplots)
#>
#> Attaching package: 'gplots'
#> The following object is masked from 'package:stats':
#>
#> lowess
library(RColorBrewer)
library(dplyr)
#>
#> Attaching package: 'dplyr'
#> The following objects are masked from 'package:stats':
#>
#> filter, lag
#> The following objects are masked from 'package:base':
#>
#> intersect, setdiff, setequal, union
library(tidyverse)
set.seed(99)
iters<-1000
#inputs
p<- 0.0005
p2<- 0.05
se<-rbeta(iters,96,6)
df<-read.csv2("df.csv")
#> Warning in file(file, "rt"): cannot open file 'df.csv': No such file or
#> directory
#> Error in file(file, "rt"): cannot open the connection
df$X<-NULL
#> Error in df$X <- NULL: object of type 'closure' is not subsettable
prp.sj<-0.1
prp.other<- 1-prp.sj
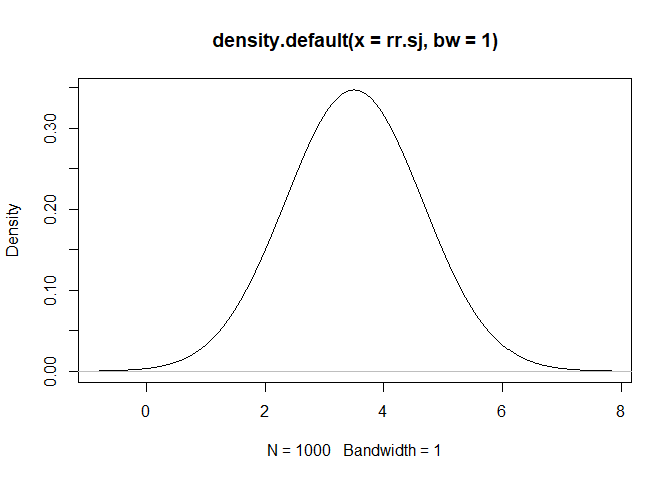
rr.sj<- rpert(iters, min = 2, mode = 3.5, max = 5)
plot(density(rr.sj, bw=1))
ar.other<-numeric(iters) #preallocate the results
ar.sj<-numeric(iters) #preallocate the results
for (i in 1:iters) {
ar.other[i]<-1/(prp.other+rr.sj[i]* prp.sj)
ar.sj[i]<-ar.other[i]*rr.sj[i]
}
prp.b<- 0.55
prp.s<- 1-prp.b
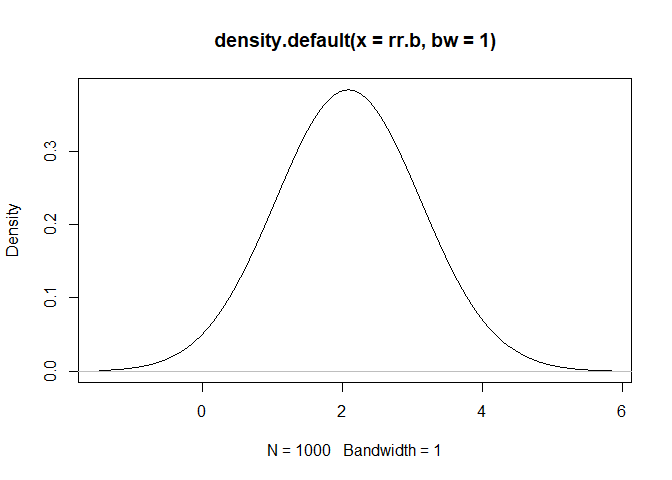
rr.b<-rpert(iters,min=1.5, mode=2, max=3)
plot(density(rr.b, bw=1))
ar.s<-numeric(iters) #preallocate the results
ar.b<-numeric(iters)#preallocate the results
for(i in 1:iters){
ar.s[i]<-1/(prp.s+rr.b[i]*prp.b)
ar.b[i]<-ar.s[i]*rr.b[i]
}
epi.h<-data.frame(other.s=numeric(iters),other.b=numeric(iters),sj.s=numeric(iters),sj.b=numeric(iters)) #preallocate dataframe
for (i in 1:iters) {
epi.h$other.s[i]<-p2*ar.other[i]*ar.s[i]
epi.h$other.b[i]<-p2*ar.other[i]*ar.b[i]
epi.h$sj.s[i]<-p2*ar.sj[i]*ar.s[i]
epi.h$sj.b[i]<-p2*ar.sj[i]*ar.b[i]
}
prp.ad<-0.2
prp.g<-1-prp.ad
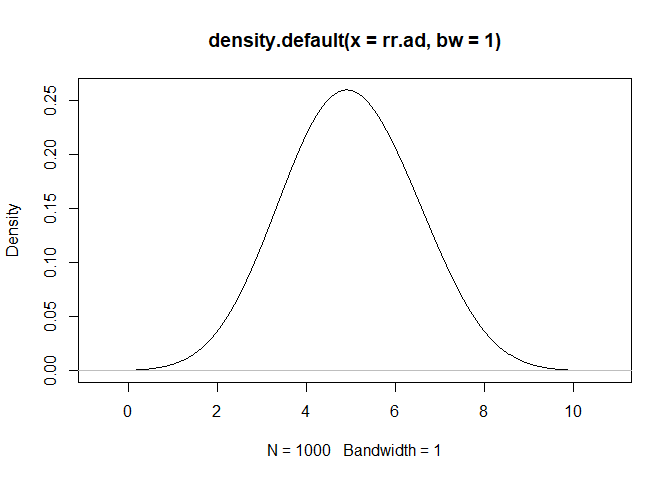
rr.ad<-rpert(iters,min=2,mode=5,max=8)
plot(density(rr.ad, bw=1))
ar.g<-numeric(iters) #preallocate the results
ar.ad<-numeric(iters)#preallocate the results
for(i in 1:iters){
ar.g[i]<-1/(prp.g+rr.ad[i]*prp.ad)
ar.ad[i]<-ar.g[i]* rr.ad[i]
}
epi.a<- data.frame(g=numeric(iters),a=numeric(iters)) #preallocate the results
for (i in 1:iters) {
epi.a$g[i]<-p*ar.g[i]
epi.a$a[i]<-p*ar.ad[i]
}
############Here the problem
#the idea is to do this operation for each month month of df$column month
m1=df %>% filter (month=="1")
#> Error in UseMethod("filter"): no applicable method for 'filter' applied to an object of class "function"
m1new<-numeric(iters)
for (i in 1:iters) { m1new[i]<- 1 - (1 - se[i] * epi.a$a[i])^m1$n[i]}
#> Error in eval(expr, envir, enclos): object 'm1' not found
##my try
results<-data.frame(m1=numeric(iters))
results<-cbind(results,rep(results[1],11))
colnames(results)<-paste("m", sep = "_", 1:12)
for (j in 1:12) {
for (i in 1:iters) {
if (df$month[i]== "1")results$m_1[i]<- 1 - (1 - se[i] * epi.a$a[i])^m1$n[i]
else if (df$month[i]== "2")results$m_2[i]<- 1 - (1 - se[i] * epi.a$a[i])^m1$n[i]
else if(df$month[i]== "3")results$m_3[i]<- 1 - (1 - se[i] * epi.a$a[i])^m1$n[i]
else if(df$month[i]== "4")results$m_4[i]<- 1 - (1 - se[i] * epi.a$a[i])^m1$n[i]
else if(df$month[i]== "5")results$m_5[i]<- 1 -(1 - se[i] * epi.a$a[i])^m1$n[i]
else if(df$month[i]== "6")results$m_6[i]<- 1 - (1 - se[i] * epi.a$a[i])^m1$n[i]
else if(df$month[i]== "7")results$m_7[i]<- 1 - (1 - se[i] * epi.a$a[i])^m1$n[i]
else if(df$month[i]== "8")results$m_8[i]<- 1 - (1 - se[i] * epi.a$a[i])^m1$n[i]
else if(df$month[i]== "9")results$m_9[i]<- 1 - (1 - se[i] * epi.a$a[i])^m1$n[i]
else if(df$month[i]== "10")results$m_10[i]<- 1 - (1 - se[i] * epi.a$a[i])^m1$n[i]
else if(df$month[i]== "11")results$m_11[i]<- 1 - (1 - se[i] * epi.a$a[i])^m1$n[i]
else if(df$month[i]== "12")results$m_12[i]<- 1 - (1 - se[i] * epi.a$a[i])^m1$n[i]
}
}
#> Error: object of type 'closure' is not subsettable
#oddly only the first column is filled.
Created on 2022-05-03 by the reprex package (v2.0.1)


