Hi everyone :D, does anyone know how I can fix this error?
Error in pts[gr, , drop = FALSE] : subscript out of bounds
library(readxl)
Family <- data.frame (tibble::tribble(
~SampleID, ~`1A`, ~`1B`, ~`1C`, ~`1D`, ~`1E`, ~`1F`, ~`1G`, ~`1H`, ~`2A`, ~`2B`, ~`2C`, ~`2D`, ~`2E`, ~`2F`, ~`2G`, ~`2H`, ~`3A`, ~`3B`, ~`3C`, ~`3D`, ~`3E`, ~`3F`, ~`3G`, ~`3H`, ~`4A`, ~`4B`, ~`4C`, ~`4D`, ~`4E`, ~`4F`, ~`4G`, ~`4H`, ~`5A`, ~`5B`, ~`5C`, ~`5D`, ~`5E`, ~`5F`, ~`5G`, ~`5H`, ~`6A`, ~`6B`, ~`6C`, ~`6D`, ~`6E`, ~`6F`, ~`6G`, ~`6H`, ~`7A`, ~`7B`, ~`7C`, ~`7D`, ~`7E`, ~`7F`, ~`7G`, ~`7H`, ~`8A`, ~`8B`, ~`8C`, ~`8D`, ~`8E`, ~`8F`, ~`8G`, ~`8H`, ~`9A`, ~`9B`, ~`9C`, ~`9D`, ~`9E`, ~`9F`, ~`9G`, ~`9H`, ~`10A`, ~`10B`, ~`10C`, ~`10D`, ~`10E`, ~`10F`, ~`10G`, ~`10H`, ~`11A`, ~`11B`, ~`11C`, ~`11D`, ~`11E`, ~`11F`, ~`11G`, ~`11H`, ~`12A`, ~`12B`, ~`12C`, ~`12D`, ~`12E`, ~`12F`, ~`12G`, ~`12H`,
"PS", 15.09523622, 15.55277853, 15.11741994, 14.554094, 15.35093463, 15.42427054, 15.95013804, 15.42427054, 15.64673681, 15.82226616, 15.89833728, 15.32111918, 15.19323748, 15.6327213, 15.42427054, 15.82409171, 15.33876286, 15.15083401, 15.66030063, 15.25702856, 15.45853597, 15.62976904, 15.42427054, 15.34240388, 15.41444138, 15.93324327, 15.97179819, 15.42427054, 15.42427054, 15.42427054, 15.14538555, 15.36670841, 14.94126609, 15.40831718, 15.0435571, 15.85328648, 15.48829559, 15.44315041, 14.91093192, 15.15380783, 15.42427054, 15.58703711, 15.60293689, 15.72780729, 15.28349865, 15.71357413, 15.42427054, 15.64458728, 14.77844862, 15.16413111, 15.42427054, 15.95385476, 15.41125872, 15.35216848, 14.77580595, 15.05406083, 15.65446864, 15.59127691, 16.0496112, 15.32687476, 15.39778217, 15.42427054, 14.80687584, 15.19724473, 15.15131767, 15.44183272, 15.07182702, 15.17657293, 15.2360694, 15.82949662, 14.86037304, 15.23637269, 15.72840703, 15.98632579, 15.3970053, 15.88057531, 15.42427054, 16.06577509, 15.20757486, 15.42427054, 14.78757775, 15.42427054, 15.03171609, 15.27112134, 15.70331648, 15.42427054, 14.85240441, 15.42861829, 16.08043349, 15.76110644, 16.12371583, 14.84752296, 15.35933445, 15.65058528, 15.18091094, 15.59099248,
"IP", 16.31972059, 16.31972059, 16.31972059, 16.31972059, 16.38068709, 16.1299595, 16.31972059, 16.23200561, 16.29262826, 16.31972059, 16.31972059, 16.31972059, 16.40060928, 16.31972059, 16.27114993, 16.31972059, 16.31972059, 16.31972059, 16.31972059, 16.43741032, 16.27004687, 16.17243104, 16.05397452, 16.31972059, 16.31972059, 16.31972059, 16.31972059, 16.1551641, 16.27912731, 16.68913472, 16.31972059, 16.31972059, 16.32830313, 16.34775976, 16.31972059, 16.31972059, 16.31972059, 16.25145003, 16.31972059, 16.36244763, 16.19753348, 16.31972059, 16.31972059, 16.31972059, 16.31972059, 16.18461024, 16.32409956, 16.31972059, 16.31972059, 16.36517663, 16.37085899, 16.31972059, 16.31972059, 16.31972059, 16.31972059, 16.31972059, 16.2969553, 16.31972059, 16.31972059, 16.31972059, 16.31972059, 16.26498461, 16.31972059, 16.31972059, 16.31972059, 16.29075471, 16.31972059, 16.31972059, 16.31972059, 16.31972059, 16.31972059, 16.31972059, 16.31972059, 16.31972059, 16.31972059, 16.31711223, 16.16311975, 16.31972059, 16.31972059, 16.32351476, 16.34158427, 16.51249303, 16.31972059, 16.31972059, 16.31972059, 16.33774505, 16.38919092, 16.31972059, 16.31972059, 16.31972059, 16.31972059, 16.31972059, 16.33387537, 16.31972059, 16.31972059, 16.31972059,
"PU", 16.20844069, 15.64870257, 16.16151234, 16.08471445, 15.87474919, 15.98531311, 15.57029977, 15.8839279, 15.87474919, 15.86551539, 15.37963, 16.29787144, 15.87474919, 15.56819076, 15.87984026, 15.64234745, 16.18458009, 16.57731921, 15.64482559, 15.87474919, 15.87474919, 15.87474919, 15.90388237, 15.90368287, 16.40390938, 15.26156221, 15.32055595, 15.8854582, 16.08375487, 15.79791277, 15.93393468, 16.08708185, 15.87474919, 15.87474919, 16.03390029, 15.4207621, 15.78047026, 15.87474919, 16.19110155, 15.87474919, 15.88727667, 15.54155664, 15.82989846, 15.47835101, 16.19476791, 15.87474919, 15.85843405, 15.64286641, 16.1840485, 15.87474919, 15.78522972, 15.4706474, 16.01464195, 16.16397874, 16.20811279, 15.88031433, 15.87474919, 15.85762583, 15.04527395, 16.34586181, 16.45360506, 16.04628641, 16.29899651, 16.16140703, 15.90886578, 15.87474919, 15.97996124, 16.22832432, 16.14372983, 15.38089157, 15.90656313, 16.57936249, 15.45892533, 14.9354136, 16.18057749, 15.87474919, 15.95654009, 15.2916829, 16.09908404, 15.86322416, 15.87474919, 15.76604825, 16.13606026, 16.27354624, 15.74515198, 15.75320903, 15.87474919, 15.7642695, 14.77429269, 15.76687342, 14.99382061, 16.17836018, 15.87474919, 15.84613629, 16.09615514, 15.8003695
)
)
Metadata <- data.frame (tibble::tribble(
~SampleID, ~SamplingPoint, ~Depth, ~Month,
"1A", "CEA1", "80 cm", "July",
"1B", "CEA2", "80 cm", "July",
"1C", "CEA4", "Interstitial", "July",
"1D", "CEA1", "80 cm", "August",
"1E", "CEA3", "80 cm", "August",
"1F", "CEA5", "80 cm", "August",
"1G", "CEA2", "80 cm", "September",
"1H", "CEA4", "80 cm", "September",
"2A", "CEA1", "80 cm", "July",
"2B", "CEA2", "Interstitial", "July",
"2C", "CEA4", "Interstitial", "July",
"2D", "CEA1", "80 cm", "August",
"2E", "CEA3", "80 cm", "August",
"2F", "CEA5", "80 cm", "August",
"2G", "CEA2", "80 cm", "September",
"2H", "CEA4", "80 cm", "September",
"3A", "CEA1", "Interstitial", "July",
"3B", "CEA2", "Interstitial", "July",
"3C", "CEA4", "80 cm", "July",
"3D", "CEA1", "Interstitial", "August",
"3E", "CEA3", "Interstitial", "August",
"3F", "CEA5", "Interstitial", "August",
"3G", "CEA2", "Interstitial", "September",
"3H", "CEA4", "Interstitial", "September",
"4A", "CEA1", "Interstitial", "July",
"4B", "CEA3", "80 cm", "July",
"4C", "CEA4", "Interstitial", "July",
"4D", "CEA1", "Interstitial", "August",
"4E", "CEA3", "Interstitial", "August",
"4F", "CEA5", "Interstitial", "August",
"4G", "CEA2", "Interstitial", "September",
"4H", "CEA4", "Interstitial", "September",
"5A", "CEA1", "80 cm", "July",
"5B", "CEA3", "80 cm", "July",
"5C", "CEA4", "Interstitial", "July",
"5D", "CEA1", "80 cm", "August",
"5E", "CEA3", "80 cm", "August",
"5F", "CEA5", "80 cm", "August",
"5G", "CEA2", "80 cm", "September",
"5H", "CEA4", "80 cm", "September",
"6A", "CEA1", "80 cm", "July",
"6B", "CEA3", "Interstitial", "July",
"6C", "CEA5", "80 cm", "July",
"6D", "CEA1", "80 cm", "August",
"6E", "CEA3", "Interstitial", "August",
"6F", "CEA5", "Interstitial", "August",
"6G", "CEA2", "Interstitial", "September",
"6H", "CEA4", "Interstitial", "September",
"7A", "CEA1", "Interstitial", "July",
"7B", "CEA3", "Interstitial", "July",
"7C", "CEA5", "80 cm", "July",
"7D", "CEA2", "80 cm", "August",
"7E", "CEA4", "80 cm", "August",
"7F", "CEA1", "80 cm", "September",
"7G", "CEA3", "80 cm", "September",
"7H", "CEA5", "80 cm", "September",
"8A", "CEA1", "Interstitial", "July",
"8B", "CEA3", "80 cm", "July",
"8C", "CEA5", "Interstitial", "July",
"8D", "CEA2", "80 cm", "August",
"8E", "CEA4", "80 cm", "August",
"8F", "CEA1", "80 cm", "September",
"8G", "CEA3", "80 cm", "September",
"8H", "CEA5", "80 cm", "September",
"9A", "CEA2", "80 cm", "July",
"9B", "CEA3", "Interstitial", "July",
"9C", "CEA5", "Interstitial", "July",
"9D", "CEA2", "Interstitial", "August",
"9E", "CEA4", "Interstitial", "August",
"9F", "CEA1", "Interstitial", "September",
"9G", "CEA3", "Interstitial", "September",
"9H", "CEA5", "Interstitial", "September",
"10A", "CEA2", "80 cm", "July",
"10B", "CEA3", "Interstitial", "July",
"10C", "CEA5", "80 cm", "July",
"10D", "CEA2", "Interstitial", "August",
"10E", "CEA4", "Interstitial", "August",
"10F", "CEA1", "Interstitial", "September",
"10G", "CEA3", "Interstitial", "September",
"10H", "CEA5", "Interstitial", "September",
"11A", "CEA2", "Interstitial", "July",
"11B", "CEA4", "80 cm", "July",
"11C", "CEA5", "Interstitial", "July",
"11D", "CEA2", "80 cm", "August",
"11E", "CEA4", "80 cm", "August",
"11F", "CEA1", "80 cm", "September",
"11G", "CEA3", "80 cm", "September",
"11H", "CEA5", "80 cm", "September",
"12A", "CEA2", "Interstitial", "July",
"12B", "CEA4", "80 cm", "July",
"12C", "CEA1", "Interstitial", "August",
"12D", "CEA2", "Interstitial", "August",
"12E", "CEA4", "Interstitial", "August",
"12F", "CEA1", "Interstitial", "September",
"12G", "CEA3", "Interstitial", "September",
"12H", "CEA5", "Interstitial", "September"
)
)
#Family <- t(Family)
attach(Family)
Family <- Family[,-1]
rownames(Family) <- SampleID
attach(Metadata)
#> The following object is masked from Family:
#>
#> SampleID
Metadata <- Metadata[,-1]
rownames(Metadata) <- SampleID
Metadata$Month <- factor(Metadata$Month,
levels = c("July", "August", "September"))
#remove the rare microbes. It keeps only the microbes that are present in at least 10% of the samples
dim(Family)
#> [1] 3 96
Family <- Family[,colMeans(Family) >=.1]
dim(Family)
#> [1] 3 96
library(vegan)
#> Loading required package: permute
#> Loading required package: lattice
#> This is vegan 2.5-6
#NMDS
#autotransform=F evita la transformaci?n de datos
#sratmax=0.999999 para que el an?lisis no se detenga antes de tiempo cuando se tienen muchos datos
Family.mds <- metaMDS(Family, k=2, distance="bray", trymax=100, zerodist="add", autotransform=F)
#> Run 0 stress 0
#> Run 1 stress 0
#> ... Procrustes: rmse 0.1345849 max resid 0.1571215
#> Run 2 stress 0
#> ... Procrustes: rmse 0.1468844 max resid 0.1827923
#> Run 3 stress 0
#> ... Procrustes: rmse 0.1098086 max resid 0.1334504
#> Run 4 stress 0
#> ... Procrustes: rmse 0.1313742 max resid 0.1672068
#> Run 5 stress 0
#> ... Procrustes: rmse 0.09406877 max resid 0.1117154
#> Run 6 stress 0
#> ... Procrustes: rmse 0.1436202 max resid 0.1704672
#> Run 7 stress 0
#> ... Procrustes: rmse 0.1061797 max resid 0.1250528
#> Run 8 stress 0
#> ... Procrustes: rmse 0.065344 max resid 0.08156737
#> Run 9 stress 0
#> ... Procrustes: rmse 0.1522574 max resid 0.2125494
#> Run 10 stress 0
#> ... Procrustes: rmse 0.2055193 max resid 0.2714054
#> Run 11 stress 0
#> ... Procrustes: rmse 0.2100263 max resid 0.293609
#> Run 12 stress 0
#> ... Procrustes: rmse 0.1987479 max resid 0.2068764
#> Run 13 stress 0
#> ... Procrustes: rmse 0.1371842 max resid 0.1810412
#> Run 14 stress 0
#> ... Procrustes: rmse 0.1130561 max resid 0.1302699
#> Run 15 stress 0
#> ... Procrustes: rmse 0.2269855 max resid 0.2939715
#> Run 16 stress 0
#> ... Procrustes: rmse 0.191206 max resid 0.2006544
#> Run 17 stress 0
#> ... Procrustes: rmse 0.2026041 max resid 0.2111073
#> Run 18 stress 0
#> ... Procrustes: rmse 0.06649483 max resid 0.08095242
#> Run 19 stress 0
#> ... Procrustes: rmse 0.1123245 max resid 0.1296081
#> Run 20 stress 0
#> ... Procrustes: rmse 0.0402027 max resid 0.05011029
#> Run 21 stress 0
#> ... Procrustes: rmse 0.1102389 max resid 0.1280593
#> Run 22 stress 0
#> ... Procrustes: rmse 0.2595968 max resid 0.3187138
#> Run 23 stress 0
#> ... Procrustes: rmse 0.1418331 max resid 0.1635621
#> Run 24 stress 0
#> ... Procrustes: rmse 0.1657882 max resid 0.2194089
#> Run 25 stress 0
#> ... Procrustes: rmse 0.1332451 max resid 0.1496957
#> Run 26 stress 0
#> ... Procrustes: rmse 0.2096469 max resid 0.2599741
#> Run 27 stress 0
#> ... Procrustes: rmse 0.06364608 max resid 0.07761915
#> Run 28 stress 0
#> ... Procrustes: rmse 0.09055523 max resid 0.1112096
#> Run 29 stress 0
#> ... Procrustes: rmse 0.1312138 max resid 0.1494752
#> Run 30 stress 0
#> ... Procrustes: rmse 0.1309514 max resid 0.148171
#> Run 31 stress 0
#> ... Procrustes: rmse 0.05396184 max resid 0.06712616
#> Run 32 stress 0
#> ... Procrustes: rmse 0.02480955 max resid 0.03243993
#> Run 33 stress 0
#> ... Procrustes: rmse 0.1166931 max resid 0.1390499
#> Run 34 stress 0
#> ... Procrustes: rmse 0.09566429 max resid 0.1302332
#> Run 35 stress 0
#> ... Procrustes: rmse 0.1178524 max resid 0.1571025
#> Run 36 stress 0
#> ... Procrustes: rmse 0.08262255 max resid 0.1009439
#> Run 37 stress 0
#> ... Procrustes: rmse 0.1866796 max resid 0.1994117
#> Run 38 stress 0
#> ... Procrustes: rmse 0.1146671 max resid 0.1317499
#> Run 39 stress 0
#> ... Procrustes: rmse 0.244136 max resid 0.3120145
#> Run 40 stress 0
#> ... Procrustes: rmse 0.03765585 max resid 0.04873949
#> Run 41 stress 0
#> ... Procrustes: rmse 0.1375493 max resid 0.159684
#> Run 42 stress 0
#> ... Procrustes: rmse 0.04855299 max resid 0.06061591
#> Run 43 stress 0
#> ... Procrustes: rmse 0.07242498 max resid 0.09028123
#> Run 44 stress 0
#> ... Procrustes: rmse 0.185853 max resid 0.2395485
#> Run 45 stress 0
#> ... Procrustes: rmse 0.2325602 max resid 0.3048931
#> Run 46 stress 0
#> ... Procrustes: rmse 0.112971 max resid 0.136291
#> Run 47 stress 0
#> ... Procrustes: rmse 0.08997876 max resid 0.1203304
#> Run 48 stress 0
#> ... Procrustes: rmse 0.2047959 max resid 0.2613437
#> Run 49 stress 0
#> ... Procrustes: rmse 0.04408303 max resid 0.05474143
#> Run 50 stress 0
#> ... Procrustes: rmse 0.168473 max resid 0.1849309
#> Run 51 stress 0
#> ... Procrustes: rmse 0.1125476 max resid 0.1297432
#> Run 52 stress 0
#> ... Procrustes: rmse 0.1168605 max resid 0.1343475
#> Run 53 stress 0
#> ... Procrustes: rmse 0.1914945 max resid 0.2014449
#> Run 54 stress 0
#> ... Procrustes: rmse 0.09439259 max resid 0.1153946
#> Run 55 stress 0
#> ... Procrustes: rmse 0.1023114 max resid 0.1238863
#> Run 56 stress 0
#> ... Procrustes: rmse 0.2180059 max resid 0.3062894
#> Run 57 stress 0
#> ... Procrustes: rmse 0.1845164 max resid 0.1970389
#> Run 58 stress 0
#> ... Procrustes: rmse 0.05031243 max resid 0.06690046
#> Run 59 stress 0
#> ... Procrustes: rmse 0.1913063 max resid 0.2639918
#> Run 60 stress 0
#> ... Procrustes: rmse 0.0343796 max resid 0.04395151
#> Run 61 stress 0
#> ... Procrustes: rmse 0.09514829 max resid 0.1136443
#> Run 62 stress 0
#> ... Procrustes: rmse 0.1731753 max resid 0.1849838
#> Run 63 stress 0
#> ... Procrustes: rmse 0.1101407 max resid 0.1275992
#> Run 64 stress 0
#> ... Procrustes: rmse 0.08100763 max resid 0.1034807
#> Run 65 stress 0
#> ... Procrustes: rmse 0.1401588 max resid 0.1584736
#> Run 66 stress 0
#> ... Procrustes: rmse 0.1123464 max resid 0.1347515
#> Run 67 stress 0
#> ... Procrustes: rmse 0.09334535 max resid 0.1103313
#> Run 68 stress 0
#> ... Procrustes: rmse 0.05576913 max resid 0.06960781
#> Run 69 stress 0
#> ... Procrustes: rmse 0.1193046 max resid 0.1369925
#> Run 70 stress 0
#> ... Procrustes: rmse 0.1456745 max resid 0.1855471
#> Run 71 stress 0
#> ... Procrustes: rmse 0.1011683 max resid 0.1216855
#> Run 72 stress 0
#> ... Procrustes: rmse 0.1846276 max resid 0.2471383
#> Run 73 stress 0
#> ... Procrustes: rmse 0.1473411 max resid 0.1683475
#> Run 74 stress 0
#> ... Procrustes: rmse 0.07551493 max resid 0.09403339
#> Run 75 stress 0
#> ... Procrustes: rmse 0.09537604 max resid 0.1279359
#> Run 76 stress 0
#> ... Procrustes: rmse 0.1172307 max resid 0.1403762
#> Run 77 stress 0
#> ... Procrustes: rmse 0.08916962 max resid 0.1058538
#> Run 78 stress 0
#> ... Procrustes: rmse 0.06390933 max resid 0.07784681
#> Run 79 stress 0
#> ... Procrustes: rmse 0.1435814 max resid 0.1958457
#> Run 80 stress 0
#> ... Procrustes: rmse 0.1186567 max resid 0.163922
#> Run 81 stress 0
#> ... Procrustes: rmse 0.0814758 max resid 0.10752
#> Run 82 stress 0
#> ... Procrustes: rmse 0.1443218 max resid 0.1597057
#> Run 83 stress 0
#> ... Procrustes: rmse 0.1206594 max resid 0.1401694
#> Run 84 stress 0
#> ... Procrustes: rmse 0.1162707 max resid 0.1497814
#> Run 85 stress 0
#> ... Procrustes: rmse 0.1054799 max resid 0.1448007
#> Run 86 stress 0
#> ... Procrustes: rmse 0.126263 max resid 0.158324
#> Run 87 stress 0
#> ... Procrustes: rmse 0.04267561 max resid 0.05496852
#> Run 88 stress 0
#> ... Procrustes: rmse 0.1814412 max resid 0.2449821
#> Run 89 stress 0
#> ... Procrustes: rmse 0.2069519 max resid 0.2130278
#> Run 90 stress 0
#> ... Procrustes: rmse 0.1180169 max resid 0.1350293
#> Run 91 stress 0
#> ... Procrustes: rmse 0.1578048 max resid 0.1897543
#> Run 92 stress 0
#> ... Procrustes: rmse 0.170893 max resid 0.1850118
#> Run 93 stress 0
#> ... Procrustes: rmse 0.1579841 max resid 0.2099756
#> Run 94 stress 0
#> ... Procrustes: rmse 0.147012 max resid 0.182826
#> Run 95 stress 0
#> ... Procrustes: rmse 0.1566814 max resid 0.1761605
#> Run 96 stress 0
#> ... Procrustes: rmse 0.04060925 max resid 0.05121645
#> Run 97 stress 0
#> ... Procrustes: rmse 0.1164699 max resid 0.1378495
#> Run 98 stress 0
#> ... Procrustes: rmse 0.03551943 max resid 0.04579404
#> Run 99 stress 0
#> ... Procrustes: rmse 0.2135406 max resid 0.2815361
#> Run 100 stress 0
#> ... Procrustes: rmse 0.1365829 max resid 0.1553028
#> *** No convergence -- monoMDS stopping criteria:
#> 100: stress < smin
#> Warning in metaMDS(Family, k = 2, distance = "bray", trymax = 100, zerodist =
#> "add", : stress is (nearly) zero: you may have insufficient data
#> Warning in postMDS(out$points, dis, plot = max(0, plot - 1), ...): skipping
#> half-change scaling: too few points below threshold
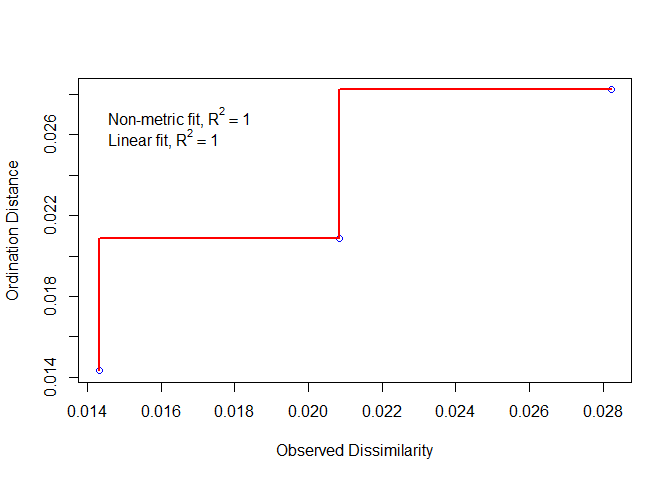
stressplot(Family.mds)
#add the metadata information
##THIS HAS TO BE IN THE SAME ORDER IN THE METADATA FILE AND IN THE COUNTS FILE
colvec<- c("yellowgreen","turquoise4", "tomato2")
plot(Family.mds, type="n", xlim=c(-.5,.5), ylim=c(-0.6,0.6))
pl <-ordiellipse(Family.mds, Metadata$Month, kind="se", conf=0.95, lwd=2, col="gray30", label=T)
#> Error in pts[gr, , drop = FALSE]: subíndice fuera de los límites
Created on 2021-07-08 by the reprex package (v0.3.0)