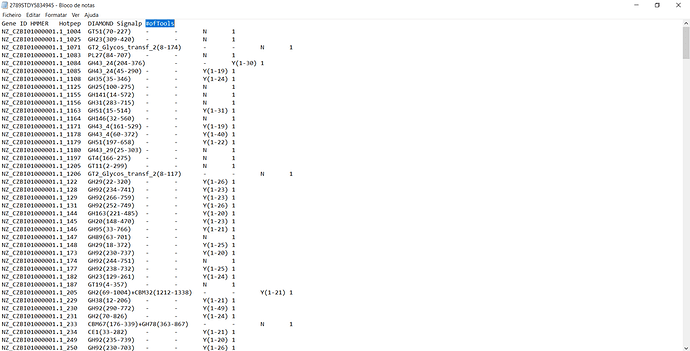
Thanks for the reply! Here it is:
> dbcan_files <- list.files("C:/Users/Lu/DBCA/",pattern="txt")
> dbcan_all <- data.frame("GH"="x","file_name"="a",stringsAsFactors=F)[-1,]
> for (f in dbcan_files) {
+ file <- read.delim(paste0("C:/Users/Lu/DBCA/",f))
+ file_name <- basename(paste0("C:/Users/Lu/DBCA/",f))
+ file_name <- gsub(".txt", "19_BTHE", file_name)
+ file_name <- gsub(".txt", "VPI-5482", file_name)
+ file_name <- gsub(".txt", "VPI-5482 (1)", file_name)
+ file_name <- gsub(".txt", "OF03-10BH", file_name)
+ file_name <- gsub(".txt", "OF05-16BH", file_name)
+ file_name <- gsub(".txt", "NCTC13706", file_name)
+ file_name <- gsub(".txt", "NCTC10582", file_name)
+ file_name <- gsub(".txt", "MGYG-HGUT-00196", file_name)
+ file_name <- gsub(".txt", "F9-2", file_name)
+ file_name <- gsub(".txt", "CL15T12C11", file_name)
+ file_name <- gsub(".txt", "bq_0049", file_name)
+ file_name <- gsub(".txt", "Bacteroides_thetaiotaomicron_81H8", file_name)
+ file_name <- gsub(".txt", "ATCC_29741", file_name)
+ file_name <- gsub(".txt", "AM51-2", file_name)
+ file_name <- gsub(".txt", "AM30-26", file_name)
+ file_name <- gsub(".txt", "AM26-17", file_name)
+ file_name <- gsub(".txt", "AM15-10", file_name)
+ file_name <- gsub(".txt", "AM09-21", file_name)
+ file_name <- gsub(".txt", "AF45-16BH", file_name)
+ file_name <- gsub(".txt", "AF28-2Y", file_name)
+ file_name <- gsub(".txt", "AF24-19LB", file_name)
+ file_name <- gsub(".txt", "AF14-20", file_name)
+ file_name <- gsub(".txt", "AF03-26", file_name)
+ file_name <- gsub(".txt", "AD135X1B", file_name)
+ file_name <- gsub(".txt", "7330", file_name)
+ file_name <- gsub(".txt", "2789STDY5834945", file_name)
+ file_name <- gsub(".txt", "2789STDY5834899", file_name)
+ file_name <- gsub(".txt", "2789STDY5834846", file_name)
+ file_name <- gsub(".txt", "2789STDY5608873", file_name)
+ file_name <- gsub(".txt", "19_BTHE", file_name)
+ file_name <- gsub(".txt", "14-106904-2", file_name)
+ names(file) <- c("gene", "hmmer", "hotpet", "diamond", "signalp", "#ofTools")
+ f_1 <- file%>%
+ filter(n_tools >= 2)%>%
+ mutate_all(as.character)
+ f_1$hmmer <-sapply(strsplit(f_1$hmmer, "[()]"), function(x) x[[1]][1])
+ f_1$hmmer <-sapply(strsplit(f_1$hmmer, "_"), function(x) x[[1]][1])
+ f_1[f_1=="N"] <- NA
+ f_2 <- f_1 %>%
+ dplyr::select(gene, hmmer)%>%
+ filter(grepl("GH", hmmer))%>%
+ group_by(hmmer)%>%
+ dplyr::summarise(n = n())
+ names(f_2) <- c("GH", file_name)
+ dbcan_all <- full_join(dbcan_all,f_2,by=c("GH"="GH"))
+ }
Error: Problem with `filter()` input `..1`.
x object 'n_tools' not found
i Input `..1` is `n_tools >= 2`.
Run `rlang::last_error()` to see where the error occurred.