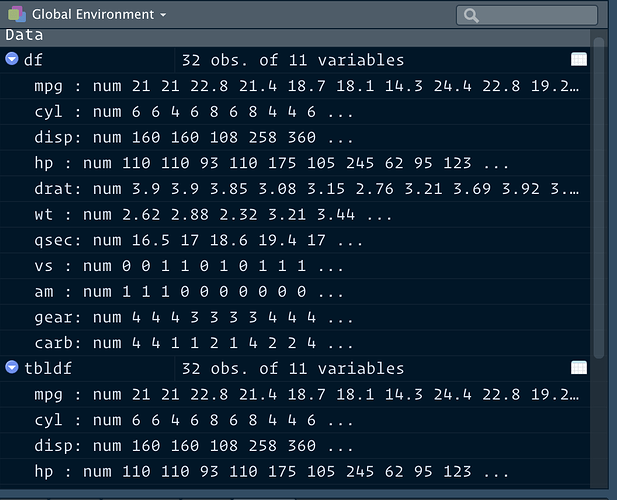
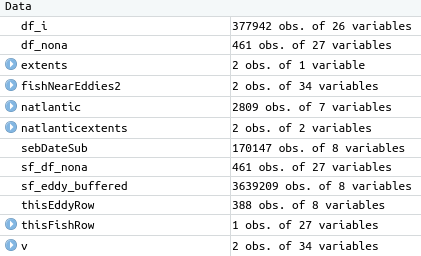
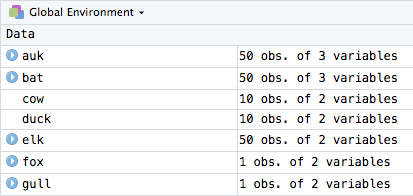
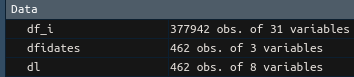
Ok, opening both original csv and saved Rds then doing all.equal() shows me that the basic read.csv() only has column types integer, and factor. The Rds also has: numeric, character, Date, and hms/difftime
So: close rstudio, open rstudio, readRdS(), object doesn't expand
Remove object, library(lubridate), readRDS(), object doesn't expand
Remove object, library(tidyverse), readRDS(), object doesn't expand
Remove object, library(dplyr), readRDS(), object doesn't expand
Delete Date column: works immediately.
Restart rstudio, no library(), readRDS(), delete date column, works.
Remove object, readRDS(), delete hms/difftime columns, still won't expand.
CULPRIT: 'Date' format columns prevent environment pane data from being expandable.
str(sharks)
$ Date : Date, format: "2006-05-15" "2006-05-15"
- attr(*, "spec")=List of 3
..$ cols :List of 79
.. ..$ Date : list()
.. .. ..- attr(*, "class")= chr "collector_character" "collector"
However, this doesn't cause any problems:
tmp <- data.frame(a = c(1,2),
Date = c("2006-05-15","2006-05-14"))
tmp$Date <- as.Date(tmp$Date)
saveRDS(object = tmp, file = "tmp.Rds")
tmp <- readRDS("tmp.Rds")
So maybe it's not that it's a Date format column specifically but it's something about the data within it?
summary(sharks$Date)
Min. 1st Qu. Median Mean 3rd Qu. Max.
"2006-05-15" "2009-03-31" "2012-05-29" "2012-07-29" "2015-09-22" "2018-11-08"
class(sharks$Date)
[1] "Date"
View(sharks)
# all looks normal
So that's me stumped. For some odd reason a Date format column seems to be causing the expando to not work even though Date columns don't necessarily cause these problems.
datecol <- sharks$Date
othercol1 <- rep(NA, 1092)
othercol2 <- rep(NA, 1092)
tmpdf <- data.frame(othercol1 = othercol1,
othercol2 = othercol2,
Date = datecol)
tmpdf <- tmpdf[, -3] #date column
tmpdf has no expando untill the date column is deleted, again suggesting it's something about the data, despite it all looking fine.