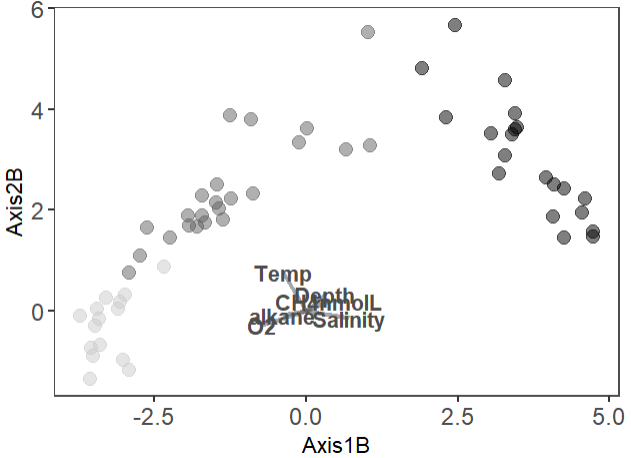
Hello, I am trying to add envfit to my NMDS plot made in ggplot and my arrows are disproportionate to the plot, where the arrows are too small.
Here is my code:
df = read.csv("df.csv", header = TRUE)
com = df[,10:11] #where columns 10 and 11 are my NMDS scores
env = pc[,2:9] #where columns 2:9 are my environmental variables
en = envfit(com, env, permutations = 9999, na.rm = TRUE)
en
en_coord_cont = as.data.frame(scores(en, "vectors")) * ordiArrowMul(en)
en_coord_cat = as.data.frame(scores(en, "factors")) * ordiArrowMul(en)
com$Cluster = pc$Cluster
gg = ggplot(data = com, aes(x = Axis1B, y = Axis2B)) +
geom_point(data = com, aes(colour = Cluster), size = 3, alpha = 0.5) +
scale_colour_manual(values = c("#CBCBCB", "#616161", "#000000")) +
geom_segment(aes(x = 0, y = 0, xend = Axis1B, yend = Axis2B),
data = en_coord_cont, size =1, alpha = 0.5, colour = "grey30") + geom_text(data = en_coord_cont, aes(x = Axis1B, y = Axis2B), colour = "grey30", fontface = "bold", label = row.names(en_coord_cont)) +
theme(panel.background = element_blank(),
panel.border = element_rect(fill = NA, colour = "grey30"),
axis.text = element_text(size = 12),
legend.key = element_blank(),
legend.title = element_text(size = 14, face = "bold", colour = "black"),
legend.text = element_text(size = 9, colour = "black"))
gg
I can increase the length of the arrows by multiplying by a larger integer in en_coord_cont = as.data.frame(scores(en, "vectors")) * ordiArrowMul(en) such as en_coord_cont = as.data.frame(scores(en, "vectors")) * ordiArrowMul(en) * 2 but from my understanding the appropriate length of the arrows is kept in the 'ordiArrowMul(en)' so I don't want to just multiply by a random number to get longer arrows.
I installed ggvegan as I've seen it recommended by users facing the same problem but I can't seem to figure out how it works... Since I already have all the values and envfit info extracted, am I to use the autoplot() command? Can I take my plot (object 'gg' and directly input it into ggvegan? I apologize for my lack of understanding of the package and R generally...
Any help is appreciated thank you.