Hello,
I am investigating linear models of a many different variables, so my ultimate goal is to make many scatterplots with changing variables, labels, and scales. Here is the workflow to do so (edited to do by reprex):
EDIT AGAIN: See display ggplot without updating values - #6 by Megan_Huber comment below for the correct summary of what's happening here - I was wrong about which variables were affected.
library(tidyverse)
library(cowplot)
# define a function to make plots:
gg_lm_simple <- function(.data = ., .y, .x, .var = NULL, .se = FALSE, ...) {
ggplot(.data, aes(x = {{.x}}, y = {{.y}})) +
geom_jitter(aes(fill = {{.var}}), shape = 21, na.rm = TRUE, alpha = 0.7, size = 2, ...) +
geom_smooth(method = "lm", formula = "y ~ x", se = {{.se}}, color = "black", size = 1) +
scale_fill_distiller(type = "div", palette = "RdBu",
aesthetics = c("fill", "color"), direction = 1,
limits = c(-1,1)*max(abs({{.var}}))) + #to keep center at 0, which is necessary for my data
theme_minimal() +
labs(title = bquote(Delta ~ .(outcome_oi_name) ~ "~" ~ Delta ~ .(var_oi_name)),
y = bquote(Delta ~ .(outcome_oi_name) ~ .(outcome_oi_unit)),
x = bquote(Delta ~ .(indep_oi_name) ~ .(indep_oi_unit)),
fill = bquote(Delta ~ .(var_oi_name) ~ .(var_oi_unit)))
}
df <- mtcars %>%
mutate(wt = (wt-3.2)*4) %>% #to approximate data that is both positive and negative
mutate(qsec = qsec-18) #to approximate data that is both positive and negative, smaller scale than qsec
##############################################
#And then this part I copy and paste about 48 times, updating the variables as needed:
# define which of my variables I want to use
outcome_of_interest <- df$disp
outcome_oi_name <- "Disp Name"
outcome_oi_unit <- "(disp units)"
outcome_oi_vlabel <- "disp"
indep_of_interest <- df$drat
indep_oi_name <- "Drat"
indep_oi_unit <- "(drat unit)"
indep_oi_vlabel <- "drat"
var_of_interest <- df$wt
var_oi_name <- "Weight"
var_oi_unit <- "(lbs)"
var_oi_vlabel <- "wt"
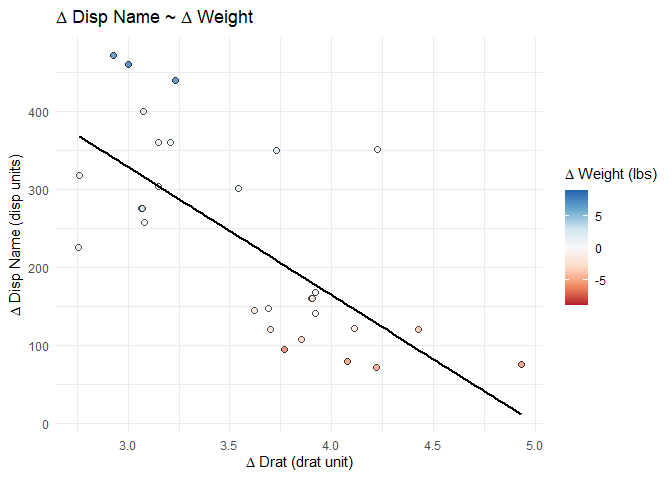
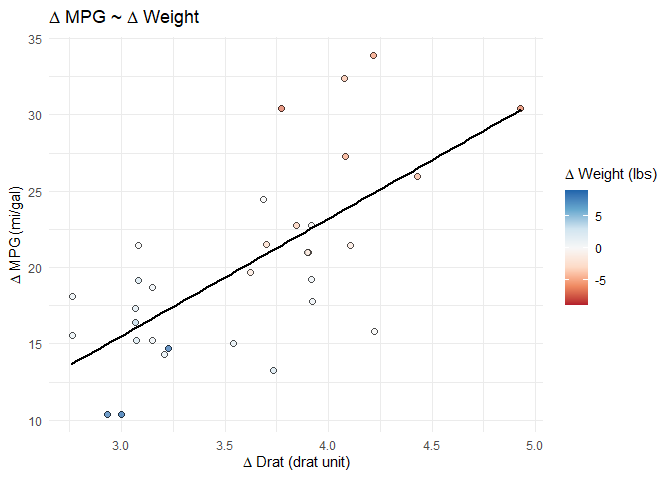
splot_out_in_var <- df %>%
gg_lm_simple(outcome_of_interest, indep_of_interest, var_of_interest)
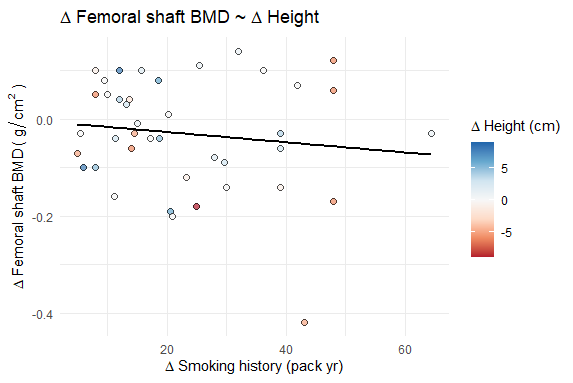
# display the plot at this point in the .Rmd
splot_out_in_var
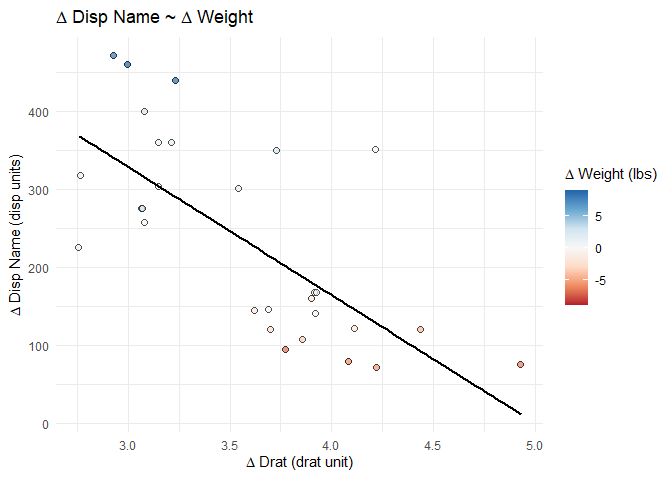
# save the plot with the specific variable names to use later
assign(glue::glue("splot_{outcome_oi_vlabel}_{indep_oi_vlabel}_{var_oi_vlabel}"), splot_out_in_var)
# so splot_out_in_var would also be assigned to splot_varA_varB_varC
# plus some other code/analysis here, which is not relevent to the question
###############################################
# new variable defined, code is copy/pasted and updated (this will be done about 47 more times)
# define which of my variables I want to use
outcome_of_interest <- df$disp
outcome_oi_name <- "Disp Name"
outcome_oi_unit <- "(disp units)"
outcome_oi_vlabel <- "disp"
indep_of_interest <- df$drat
indep_oi_name <- "Drat"
indep_oi_unit <- "(drat unit)"
indep_oi_vlabel <- "drat"
var_of_interest <- df$qsec
var_oi_name <- "Qsec"
var_oi_unit <- "(units)"
var_oi_vlabel <- "qsec"
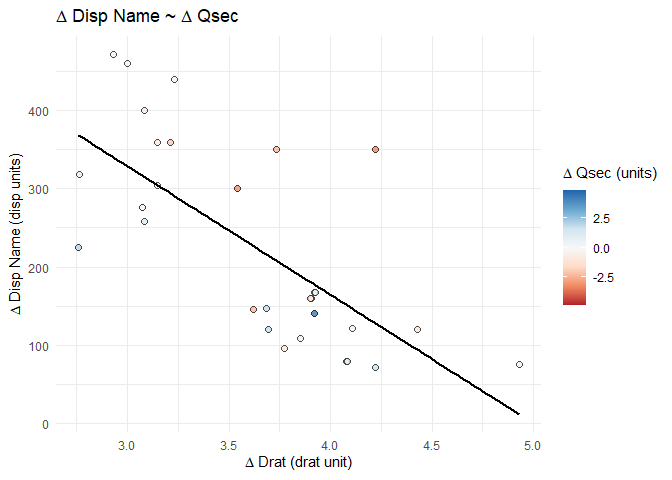
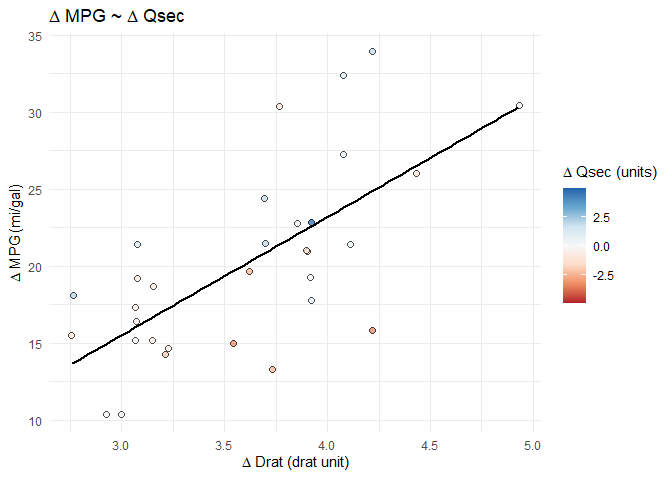
splot_out_in_var <- df %>%
gg_lm_simple(outcome_of_interest, indep_of_interest, var_of_interest)
# display the plot at this point in the .Rmd
splot_out_in_var
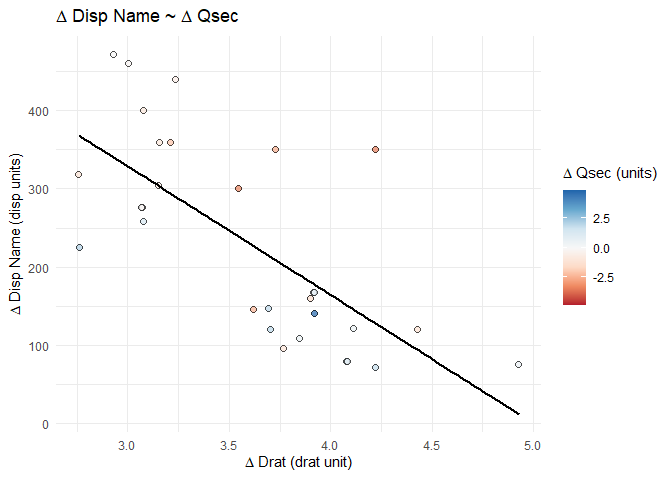
# save the plot with the specific variable names to use later
assign(glue::glue("splot_{outcome_oi_vlabel}_{indep_oi_vlabel}_{var_oi_vlabel}"), splot_out_in_var)
## so splot_out_in_var would also be assigned to splot_varA_varB_varC
max(abs(var_of_interest))
#> [1] 4.9
max(abs(df$qsec))
#> [1] 4.9
#################################################
#At the end of the Rmd, I want to compare these plots together using cowplot::plot_grid.
# Here I have specifically typed out all 48 names, I'm not using any sort of function to get the names
splot_list <- list(splot_disp_drat_wt, splot_disp_drat_qsec)
# and make the plot:
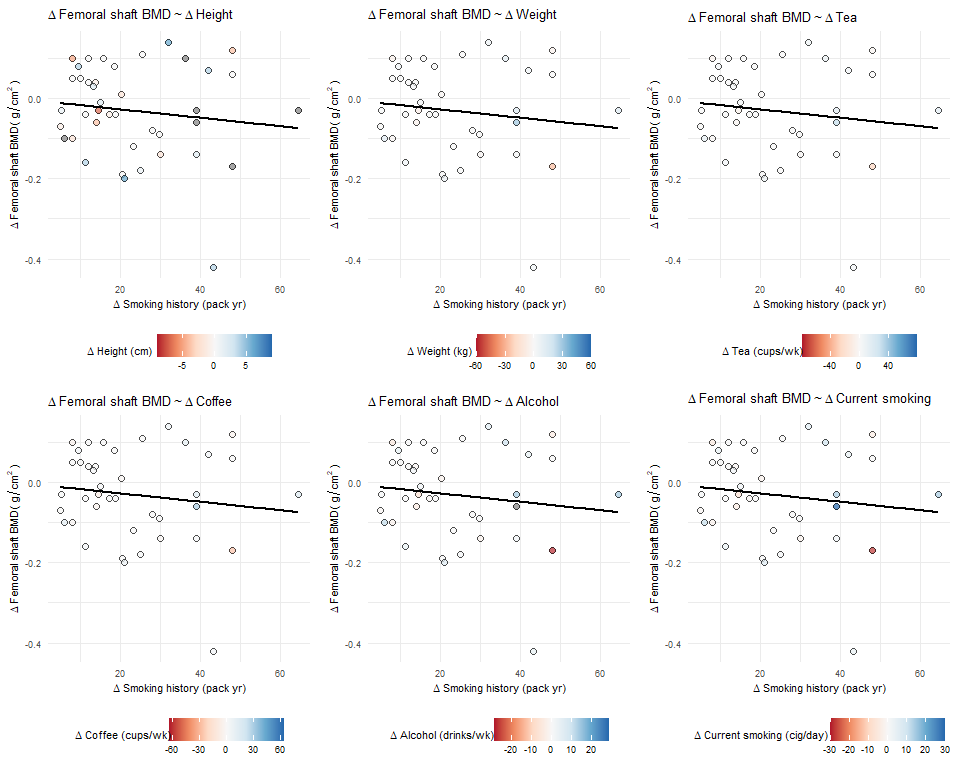
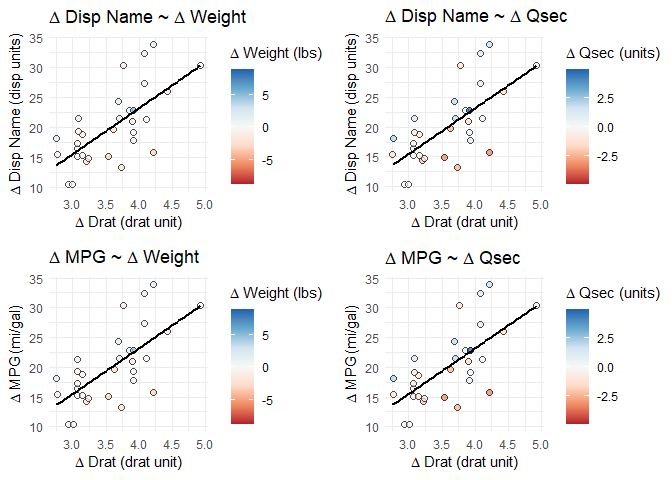
cowplot::plot_grid(plotlist = splot_list, align = "hv", nrow = 2)
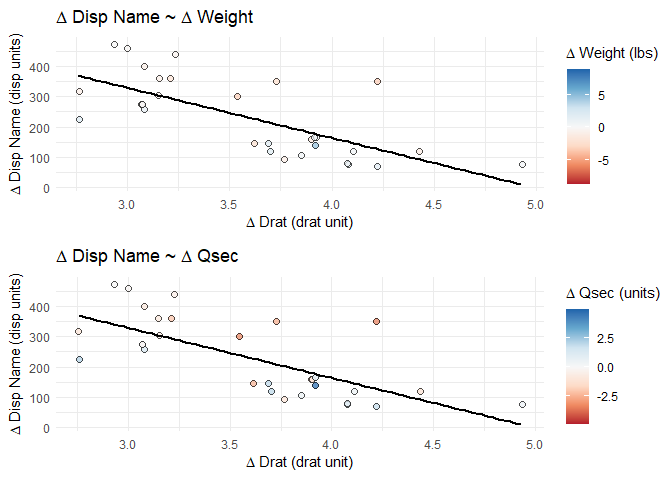
Created on 2021-03-11 by the reprex package (v1.0.0)
The final plots have the appropriate title and axis names, axis scales, and data points/regression lines. The color/fill scales show the proper values in the legend and the points have the correct relative color for the correct variable, but the actual fill scales are all based on the final plot's variable.
So for a plot that's varA~ varB + varC (where varC goes from -10,10)
When first made, the plot legend will show fill values of c(-10, 10) and the points will be filled such that a point with varC = -10 is darkest red and one with varC = 5 is medium blue, as expected.
However: I then copy/paste, change the outcome/indep/variable_of_interest, and make many new plots (so now the most recent variable_of_interest <- varZ, where varZ goes from -5,5) and call cowplot::plot_grid(). The portion of the plot with varA~ varB + varC has the appropriate titles/labels, the points are filled by varC, the fill legend still shows values from -10 to 10. But the actual scaling of fill values is dictated by varZ, not varC. So a point with varC = -5 is darkest red and one with varC = 5 is darkest blue, but a point where varC = -8 (outside the value range of varZ) is filled grey as if it has a missing varC value. The relationship between the filled points is still relative to varC and not varZ, but the fill scale is set by varZ.
I have no clue how to go from here. Is there a way to assign the ggplot to a variable name and then unlink it somehow, so that calling the variable name will not re-run the gg_lm_simple code but instead just display the saved ggplot object? Or is there something wrong with the way I designed my function so that only the values argument of scale_fill_distiller will be incorrect but everything else is fine?
I guess I could run ggsave() on each plot, then use something to load and grid each image file, but it already takes over 10 minutes to knit and my computer has limited RAM to process loading several hundred images.