Hello community,
I am new to R, so please forgive me if the answer to my question is obvious or if my code is rubbish.
I have data that represents time (on x) and a percentage (on y). "y" can thus not be higher than 100. I would like to calculate a log trendline/fitting linear model. I used the following code:
x <- c(3, 5, 7, 14, 21, 28, 56, 100, 200)
y <- c(20, 30, 50, 70, 89, 95, 99.9, 99.99, 99.999)
fitlog <-lm(y~log(x))
xx <- seq(1,200, length=200)
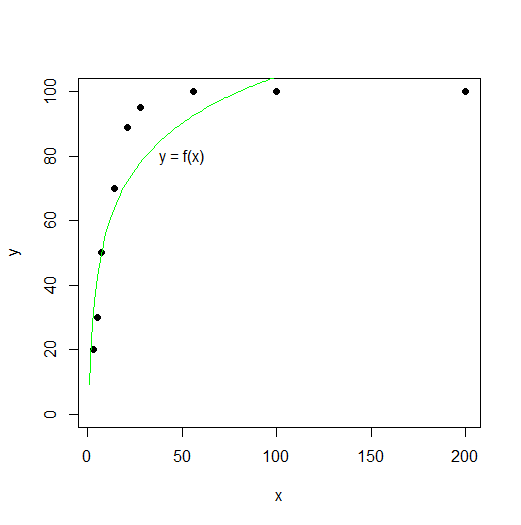
plot(x,y,pch=19,ylim=c(0,200))
lines(xx, predict(fitlog, data.frame(x=xx)), col="green")
#this way the curve goes higher than 100 (apart from also not fitting very well). I have three questions related to this project:
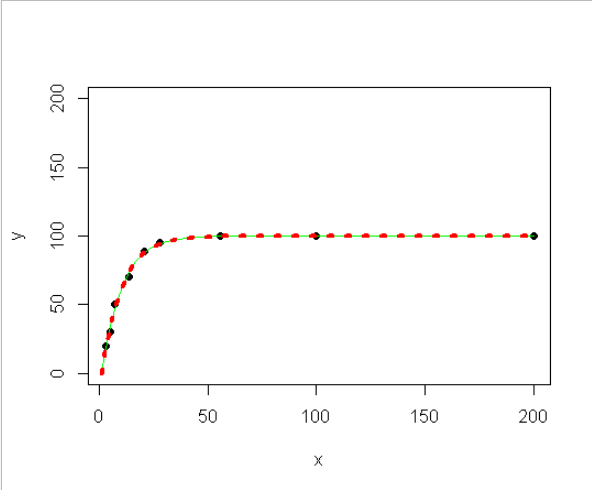
- How can I define the "y" range to be between 0-100, so that I still get the best linear modelling within this boundary?
- How can I make R display the function behind the trendline?
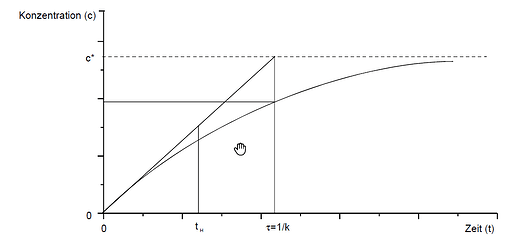
- How can I calculate the asymptote (in a similar case, where it might not be 100) with R?
Thank you very much in advance!
(I am using RStudio Version 1.2.5033 on a Windows 10 Pro 64-Bit.)
Best regards
FL