Dear all,
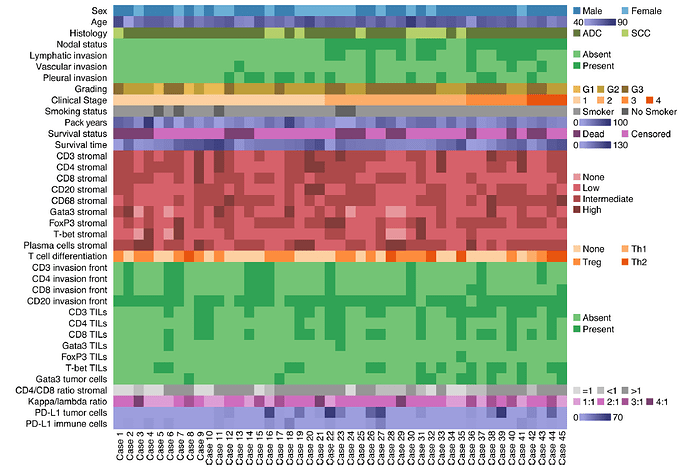
how ist it possible to create this/or a similar heatmap-like table (see below) using the following data set
Does converting the character variables to numeric ones help in the first place?
Case sex Age type subtype localisation
1 1 w 68 IPNB pancreatobiliary distal
2 2 m 62 IPNB pancreatobiliary distal
3 3 w 53 IPNB intestinal intrahepatic
4 4 m 59 IPNB pancreatobiliary distal
5 5 w 55 ITPN pancreatobiliary intrahepatic
6 6 m 69 ITPN pancreatobiliary intrahepatic
Thank you so much for taking your time to help this VERY desperate person right now...