H All,
I would like to calculate correlations between variables: P18, P19, P20, P21, P22, P23 - each variable in turn with variables: P35, P36, P37, P38, P39, P40, P41.
So first will be correlations between variable P18 and P35, P36, P37, P38, P39, P40, P41 variables and then P19 and P35, P36, P37, P38, P39, P40, P41 variables and so on.
I would like to store the results in dataframe with "correlation result" and "p-value" as columns.
How do I do it with for loop, please ? Any ideas will be greatly appreciated.
I have done it sort of manually one by one steps, but I want to do it using for loop in a more automatic way.
I have done this using manual calculations and with rbind() function later on.
I would like to do it for all rows in data frame (as a whole) and for "Location" levels subgroups ("Group1", "Group2", "Group3", "Group4") separately.
How do I do it please.
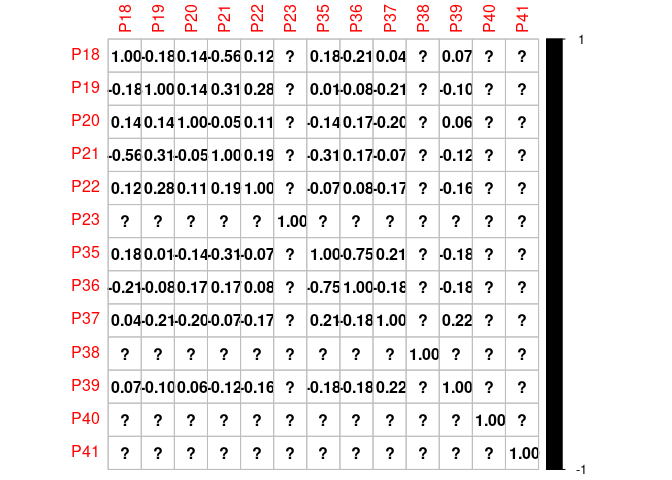
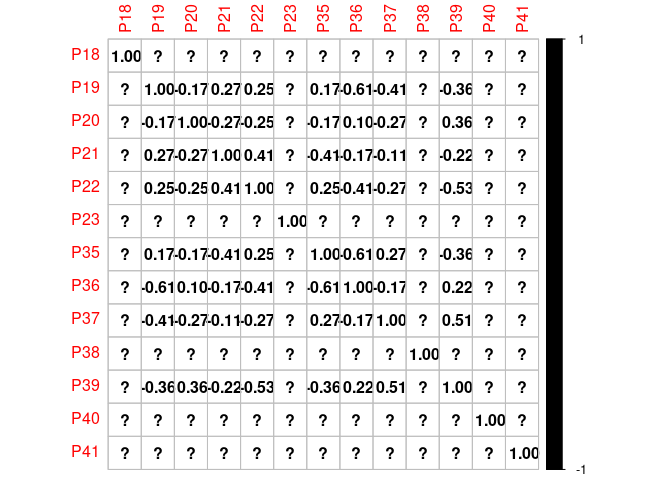
Here below there are my desired results as an example:
Here below there is my data frame:
dfM <- structure(list(Location = structure(c(1L, 1L, 1L, 1L, 1L, 1L,
1L, 1L, 1L, 1L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 3L, 3L,
3L, 3L, 3L, 3L, 3L, 3L, 3L, 3L, 4L, 4L, 4L, 4L, 4L, 4L, 4L, 4L,
4L, 4L), .Label = c("Group1", "Group2", "Group3", "Group4"), class = "factor"),
P18 = c(1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1,
1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 0, 1, 1, 1, 1, 1, 1, 1,
1, 1, 1, 1), P19 = c(1, 1, 1, 1, 1, 0, 0, 0, 0, 1, 0, 0,
0, 1, 1, 1, 1, 0, 0, 1, 0, 1, 0, 1, 0, 0, 1, 1, 1, 0, 0,
1, 0, 1, 0, 0, 0, 0, 0, 0), P20 = c(0, 0, 0, 1, 0, 1, 0,
0, 1, 1, 0, 0, 1, 1, 1, 0, 1, 0, 0, 1, 0, 1, 1, 0, 0, 1,
0, 0, 0, 0, 0, 1, 0, 1, 0, 1, 0, 1, 0, 1), P21 = c(0, 0,
1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0,
0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0),
P22 = c(0, 1, 1, 0, 1, 0, 0, 0, 1, 0, 0, 0, 0, 1, 0, 0, 0,
0, 0, 1, 1, 0, 1, 1, 0, 1, 1, 0, 0, 0, 0, 1, 1, 1, 0, 0,
0, 0, 0, 0), P23 = c(0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0), P35 = c(1, 1, 0, 0, 1, 0, 1,
0, 1, 1, 1, 1, 0, 1, 1, 1, 0, 1, 0, 0, 1, 0, 0, 0, 0, 0,
1, 1, 0, 1, 1, 1, 0, 0, 0, 1, 1, 1, 0, 1), P36 = c(0, 0,
0, 0, 0, 1, 0, 1, 0, 0, 0, 0, 1, 0, 1, 0, 0, 0, 1, 1, 0,
1, 1, 1, 1, 1, 0, 0, 1, 0, 0, 0, 1, 1, 1, 0, 0, 0, 0, 0),
P37 = c(0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0), P38 = c(0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0), P39 = c(0, 0, 0, 1, 0, 1, 1,
0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 1, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0), P40 = c(0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0),
P41 = c(0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0)), class = "data.frame", row.names = c(NA, -40L
))
thank you and kind regards.