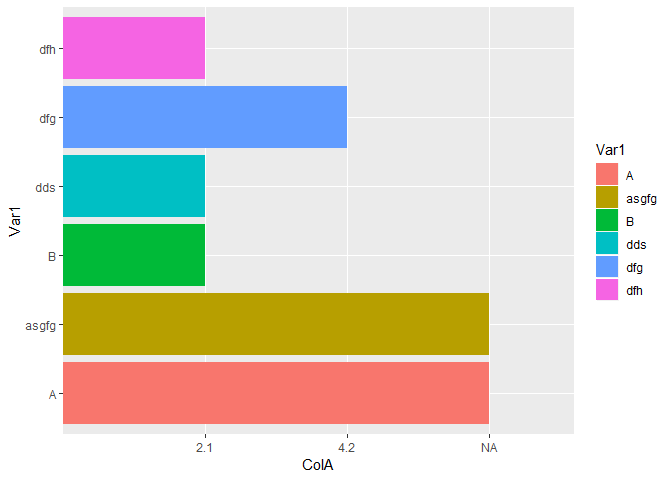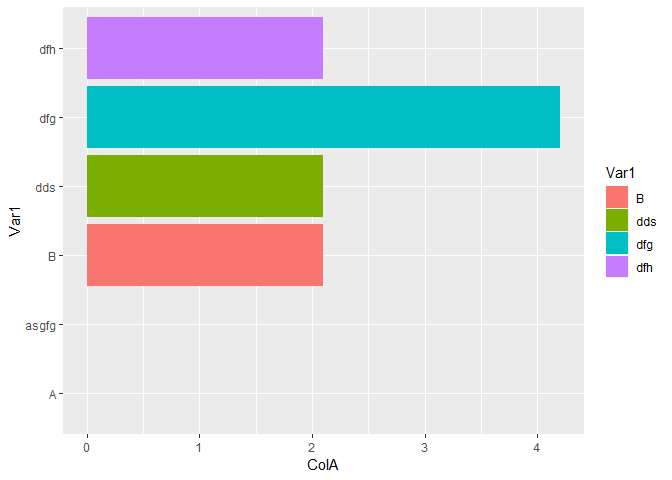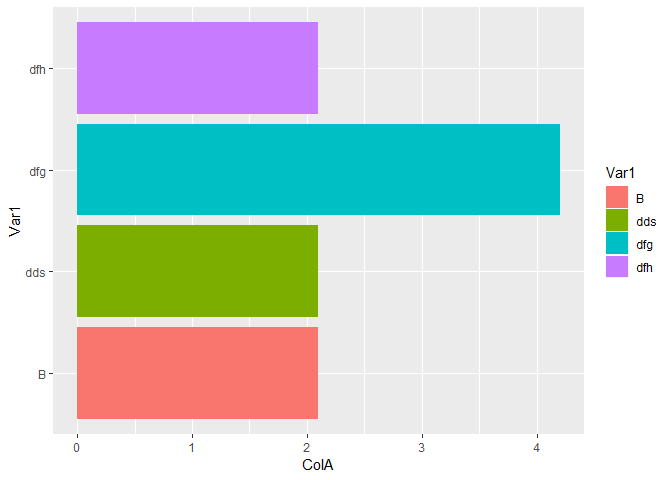Your example happens to be very confusing because the NAs in ColA are actually strings that happen to have the two letters NA. This causes the entire column to be interpreted as a factor. There is further confusion because the other two values in the column would differ by a factor of two if they were interpreted as numbers, so they look plausibly spaced on the axis. Examine these variations of your data.
library(ggplot2)
#Original
p12 <- data.frame(Var1 = c("A","asgfg","B","dds","dfg","dfh"),
ColA=c("NA","NA",2.1,2.1,4.2,2.1))
ggplot(data=p12,aes(x=Var1,y=ColA,fill=Var1))+geom_bar(stat = "identity")+coord_flip()
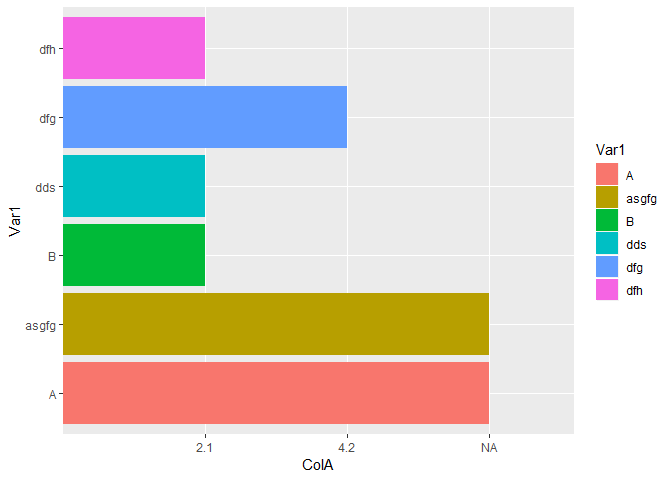
#Actual NA
p12_2 <- data.frame(Var1 = c("A","asgfg","B","dds","dfg","dfh"),
ColA=c(NA,NA,2.1,2.1,4.2,2.1))
ggplot(data=p12_2,aes(x=Var1,y=ColA,fill=Var1))+geom_bar(stat = "identity")+coord_flip()
#> Warning: Removed 2 rows containing missing values (position_stack).
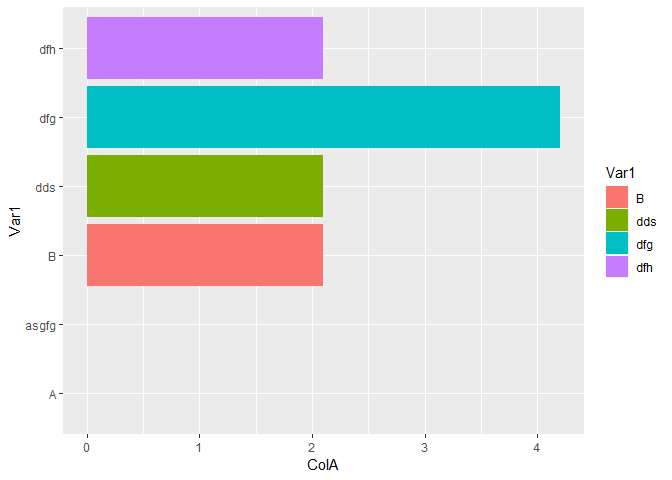
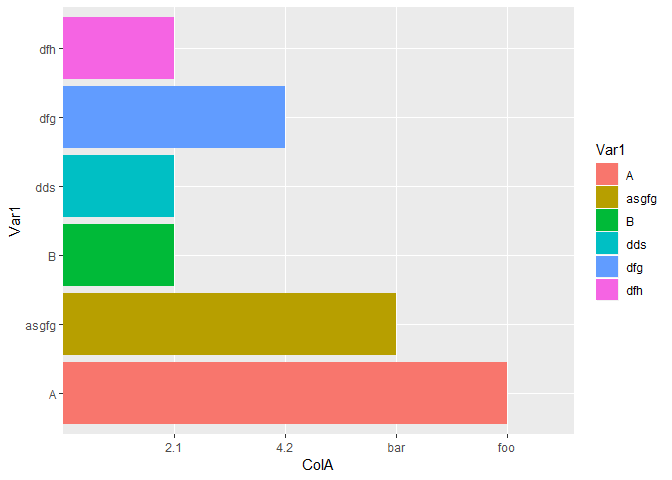
#Strings that are not "NA"
p12_3 <- data.frame(Var1 = c("A","asgfg","B","dds","dfg","dfh"),
ColA=c("foo","bar",2.1,2.1,4.2,2.1))
ggplot(data=p12_3,aes(x=Var1,y=ColA,fill=Var1))+geom_bar(stat = "identity")+coord_flip()
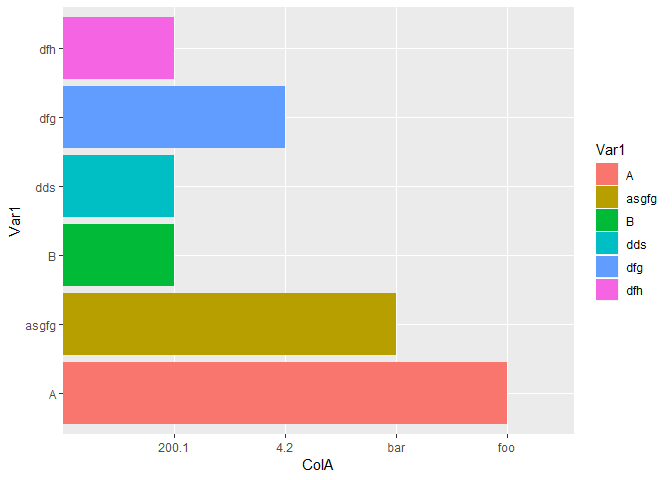
#Different ColA values
p12_4 <- data.frame(Var1 = c("A","asgfg","B","dds","dfg","dfh"),
ColA=c("foo","bar",200.1,200.1,4.2,200.1))
ggplot(data=p12_4,aes(x=Var1,y=ColA,fill=Var1))+geom_bar(stat = "identity")+coord_flip()
Created on 2019-10-01 by the reprex package (v0.2.1)
In your actual data, you probably want the text NA to be interpreted as an NA, not as a string. How are the data being entered?