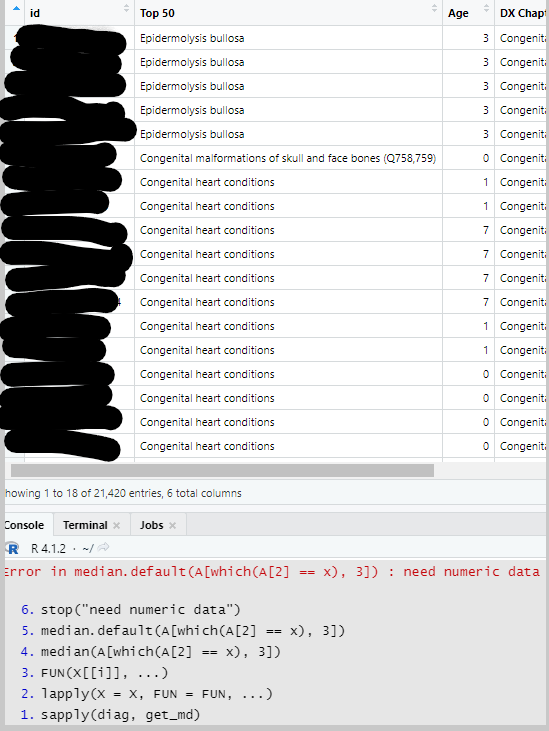
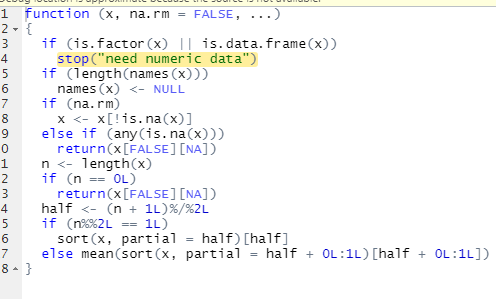
I have a dataset and am considering three of 20 variables in order to calculate median values. The three variables are id, Top 50, and Age. An id can have only one Age but it can have more than one observation from Top 50 attached to it because there are many Top 50 'types'. I need to find the median Age associated with each Top 50 observation 'type' which means I need the median Age of the distinct count of id associated with a given Top 50 observation 'type'.
I tried to create a replicable df that you all could use as an example but I CANT SEEM TO MAKE IT WORK IN RSTUDIO (super rookie over here, obv) but I do have that "df creation attempt" below so that you have an idea of the type of variables and observations that I am working with:
df<-data.frame(id = c("AXT123,"AXR456","AXV678","AXC789","AXB123","AXT124","AXV345","AXC890","AXC123","AXC345","AXV643","AXQ876","AXW345","AXE987","AXY678","AXT098","AXE345","AXT945","AXN267","AXP345"),(Top50 = c("Congenital heart conditions","Abdominal Pain","Chest Pain", "Cough","Viral infection","Cough","Cough","Viral infection","Dehydration","Abdominal Pain","Congenital heart conditions","Diabetes","Diabetes","Infection","Infection","Infection","Infection","Infection","Cancer","Congenital heart conditions"),Age = c("5","54","23","44","12","15","83","18","22","61","2","55","8","37","71","11","1","1","28","3"))
Let me know what else you need - I think I am just not considering the correct layering of commands for group_by and median but just really need some guidance.