Hi,
I want to reorder (in descending order) iris df columns that are only numerical by their means, leaving Species column intact but placing it as a first column of a df. How do I do it ?
library(tidyverse)
(what_to_do <- summarise(iris,
across(where(is.numeric),
.fns = mean)) |>
pivot_longer(cols = everything()) |>
arrange(desc(value)))
iris |> arrange_at(pull(what_to_do,name),
.funs = ~desc(.)) |> relocate(Species)Thank you,
How to after (above written code) arrange numeric columns of Iris without doing pivot_longer, please ?
So, I have got as a result of that forementioned code:
![]()
and the columns should be sorted in descending order like this:

Sepal.Length, Petal.Length, Sepal.Width, Petal.Width and as a first column I would like to have Species variable, please.
Or it could be with the usage of what_to_do object but with the result I would like to achieve as described above, thank you.
(ordered_names <- summarise(iris,
across(everything(),
.fns = mean)) |>
unlist() |>
sort(decreasing = TRUE,na.last=FALSE))Thank you very much indeed for taking time to help me.
We are almost there. I would like to have Iris dataframe sorted in decreasing order of numeric columns (according to their means), but columns should look as in original Iris dataset. With your code kindly provided I received only this:
My desired result would be that:
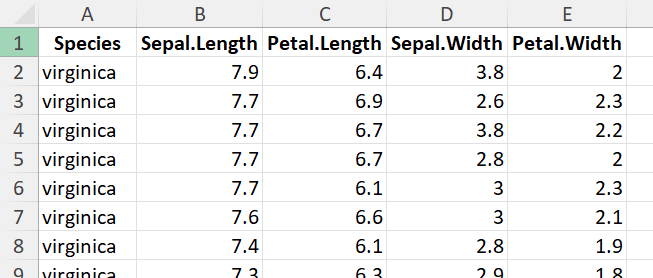
My apologies for not being clear enough from the beginning.
library(tidyverse)
(ordered_names <- summarise(iris,
across(everything(),
.fns = mean)) |>
unlist() |>
sort(decreasing = TRUE,na.last=FALSE))
(ordnm <- names(ordered_names))
iris |> arrange_at(ordnm,
.funs = ~desc(.)) |> select(all_of(ordnm))I am getting an error:
Error in `arrange_at()`:
! Can't subset columns that don't exist.
✖ Column `sort(...)` doesn't exist.
Run `rlang::last_error()` to see where the error occurred.
maybe your R session has spoiled and you could restart it ?
otherwise check the contents of ordnm; it shouldnt mention sort....
> (ordnm <- names(ordered_names))
[1] "Species" "Sepal.Length" "Petal.Length" "Sepal.Width" "Petal.Width"I did restart and cleaned environment, but in spite of that:
library(tidyverse)
ordered_names <- dplyr::summarise(iris,
across(everything(),
.fns = mean) |>
unlist() |>
sort(decreasing = TRUE, na.last = FALSE))
(ordnm <- names(ordered_names))
#> [1] "sort(...)"
iris |> arrange_at(ordnm,
.funs = ~desc(.)) |> select(all_of(ordnm))
#> Error in `arrange_at()`:
#> ! Can't subset columns that don't exist.
#> ✖ Column `sort(...)` doesn't exist.
#> Backtrace:
#> ▆
#> 1. ├─dplyr::select(arrange_at(iris, ordnm, .funs = ~desc(.)), all_of(ordnm))
#> 2. ├─dplyr::arrange_at(iris, ordnm, .funs = ~desc(.))
#> 3. │ └─dplyr:::manip_at(...)
#> 4. │ └─dplyr:::tbl_at_syms(...)
#> 5. │ └─dplyr:::tbl_at_vars(tbl, vars, .include_group_vars = .include_group_vars, error_call = error_call)
#> 6. │ ├─dplyr:::fix_call(...)
#> 7. │ │ └─base::withCallingHandlers(...)
#> 8. │ └─tidyselect::vars_select(tibble_vars, !!!vars)
#> 9. │ └─tidyselect:::eval_select_impl(...)
#> 10. │ ├─tidyselect:::with_subscript_errors(...)
#> 11. │ │ └─rlang::try_fetch(...)
#> 12. │ │ └─base::withCallingHandlers(...)
#> 13. │ └─tidyselect:::vars_select_eval(...)
#> 14. │ └─tidyselect:::walk_data_tree(expr, data_mask, context_mask)
#> 15. │ └─tidyselect:::eval_c(expr, data_mask, context_mask)
#> 16. │ └─tidyselect:::reduce_sels(node, data_mask, context_mask, init = init)
#> 17. │ └─tidyselect:::walk_data_tree(new, data_mask, context_mask)
#> 18. │ └─tidyselect:::as_indices_sel_impl(...)
#> 19. │ └─tidyselect:::as_indices_impl(...)
#> 20. │ └─tidyselect:::chr_as_locations(x, vars, call = call, arg = arg)
#> 21. │ └─vctrs::vec_as_location(...)
#> 22. └─vctrs (local) `<fn>`()
#> 23. └─vctrs:::stop_subscript_oob(...)
#> 24. └─vctrs:::stop_subscript(...)
#> 25. └─rlang::abort(...)
Created on 2022-12-13 with reprex v2.0.2
your brackets are off; the first |> after summarise has to have sufficient ) before it to close/complete the summarise action.
My silly mistake, I am truly sorry.
I would like to ask additional question, I want to adapt this code to my other df and sort it in ascending row order. I have got "items" column with items from p1 to p11. When I do it
df_other %>% arrange(items)
I got it in the following order: p1, p10, p11, p2, p3, p4, p5, p6, p7, p8, p9.
What should I do to arrange it in a proper way ? I mean: p1 , p2, p3, and so on to p11.
I think the easiest way is using readr::parse_number, as its an easy way to look past the P's and just get the numbers.
(pfalsevals <- sort(paste0("P",1:20)))
(pnums <- readr::parse_number(pfalsevals))
names(pnums) <- pfalsevals
names(sort(pnums))Thank you very much indeed, again I have learnt a lot today including how to pay attention to the smallest details such as brackets as well.
Much obliged.
This topic was automatically closed 7 days after the last reply. New replies are no longer allowed.
If you have a query related to it or one of the replies, start a new topic and refer back with a link.