The reprex below shows how to illustrate a question to attract more answers. See the FAQ: How to do a minimal reproducible example reprex for beginners.
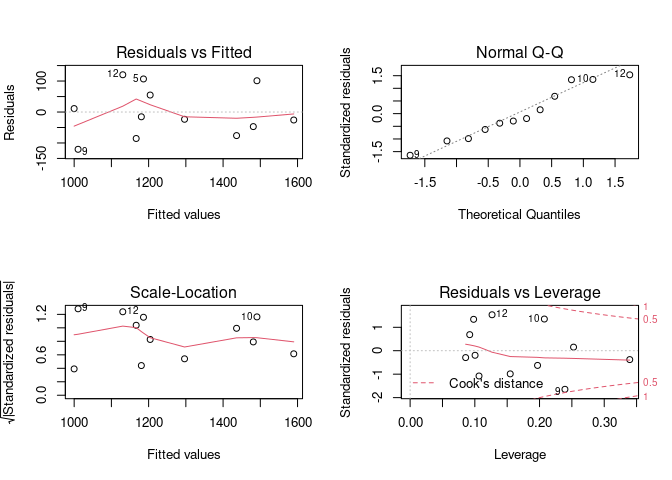
As a general rule, a data point shouldn't be excluded solely on the basis that it is at the extremes of the range unless there is a sound basis to believe that it represents an outright error—for example a pH value recorded as 16. And indeed the diagnostic plots of an lm model do not show residual outliers or highly influential points.
For the data set imaged, H6 does not appear to be an outlier either before or after treatment. Certainly not compared to H11 and H9. In fact all values are within the conventional 1.75 * interquartile distance.
dta <- data.frame(
condigo = c("H1", "H2", "H3", "H4", "H5", "H6", "H7", "H8", "H9", "H10", "H11", "H12"),
iggantes = c(1564, 1434, 1259, 1165, 1293, 1081, 1273, 1360, 890, 1592, 1011, 1251),
iggdespues = c(1705, 1572, 1233, 1204, 1211, 1187, 1346, 1517, 996, 1584, 983, 1143)
)
summary(dta)
#> condigo iggantes iggdespues
#> Length:12 Min. : 890 Min. : 983
#> Class :character 1st Qu.:1144 1st Qu.:1176
#> Mode :character Median :1266 Median :1222
#> Mean :1264 Mean :1307
#> 3rd Qu.:1378 3rd Qu.:1531
#> Max. :1592 Max. :1705
par(mfrow = c(1,2))
boxplot(dta$iggantes)
boxplot(dta$iggdespues)

fit <- lm(iggantes ~ iggdespues, data = dta)
par(mfrow = c(2,2))
plot(fit)